Our January issue is now online now open access! This issue contains 23 articles about the latest methods in ecology and evolution, including a special feature on Large data and complex models, methods on food web visualisation, biologging and much more! Read our first open access issue to find out about this month’s featured articles and the article behind our cover.
Special feature
Realising the Promise of Large Data and Complex Models
This issue includes 13 special feature articles on the topic of Large Data and Complex Models covering the exciting ways forward that technological advances allow for ecological data to be collected.
Featured Articles
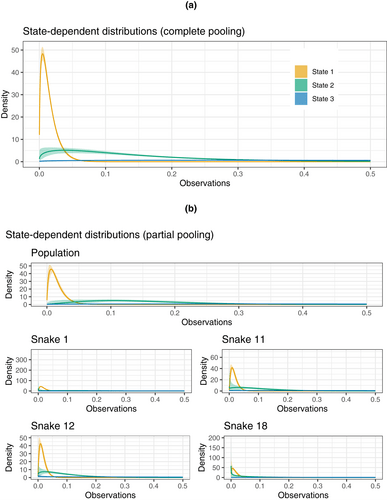
Hidden Markov models (HMMs) and their extensions are attractive methods for analysing ecological data where noisy, multivariate measurements are made of a hidden, ecological process, and where this hidden process is represented by a sequence of discrete states. This article by Grenié et al. reviews five lesser known pitfalls one can encounter when using HMMs or their extensions to solve ecological problems in order to heighten awareness of the pitfalls ecologists may encounter when applying these more advanced methods.
Harmonizing taxon names in biodiversity data: In this review Glennie et al. discuss taxonomic harmonization and the common pitfalls which can arise. The process of standardizing taxon names, taxonomic name harmonization, is necessary to properly merge data indexed by taxon names. The large variety of taxonomic databases and related tools are often not well described. As a result, taxonomic harmonization has become a major obstacle in ecological studies that seek to combine multiple datasets. In this review a set of major publicly available taxonomic databases are categorize as well as a large collection of R packages to access them and to harmonize lists of taxon names.
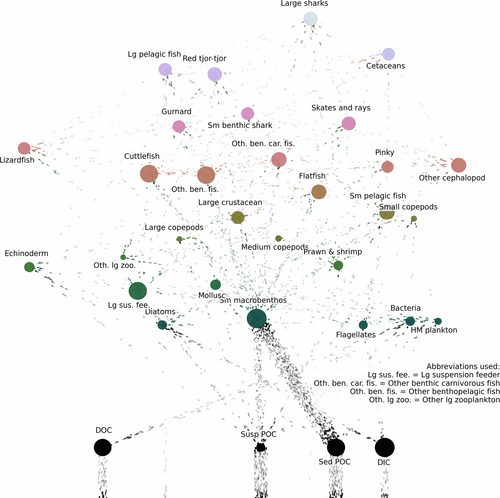
Food webs are the physical foundations of ecosystems. Visualisations help to find and present structural patterns of these weighted networks and are essential in research, conservation practice and education. This application brings several complementary and customisable methods together to facilitate accompanying every food web publication and analysis with its visualisation. Users can tinker with the interactive output and method parameters to reach the desired effect. Aesthetically appealing images presenting empirical data help to communicate the importance of species interconnectedness and ecosystem complexity to the broader public.
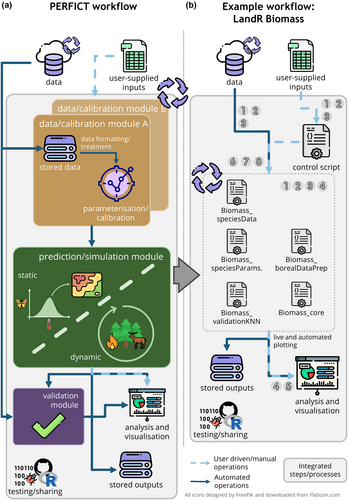
Modelling is widely used in ecology and its utility continues to increase as scientists, managers and policy-makers face pressure to effectively manage ecosystems and meet conservation goals with limited resources. As the urgency to forecast ecosystem responses to global change grows, so do the number and complexity of predictive ecological models and the value of iterative prediction, both of which demand validation and cross-model comparisons. By linking all stages of an ecological modelling exercise, it is possible to overcome common challenges faced by ecological modellers, such as changing study areas, choosing between different modelling approaches, and evaluating the appropriateness of the model.
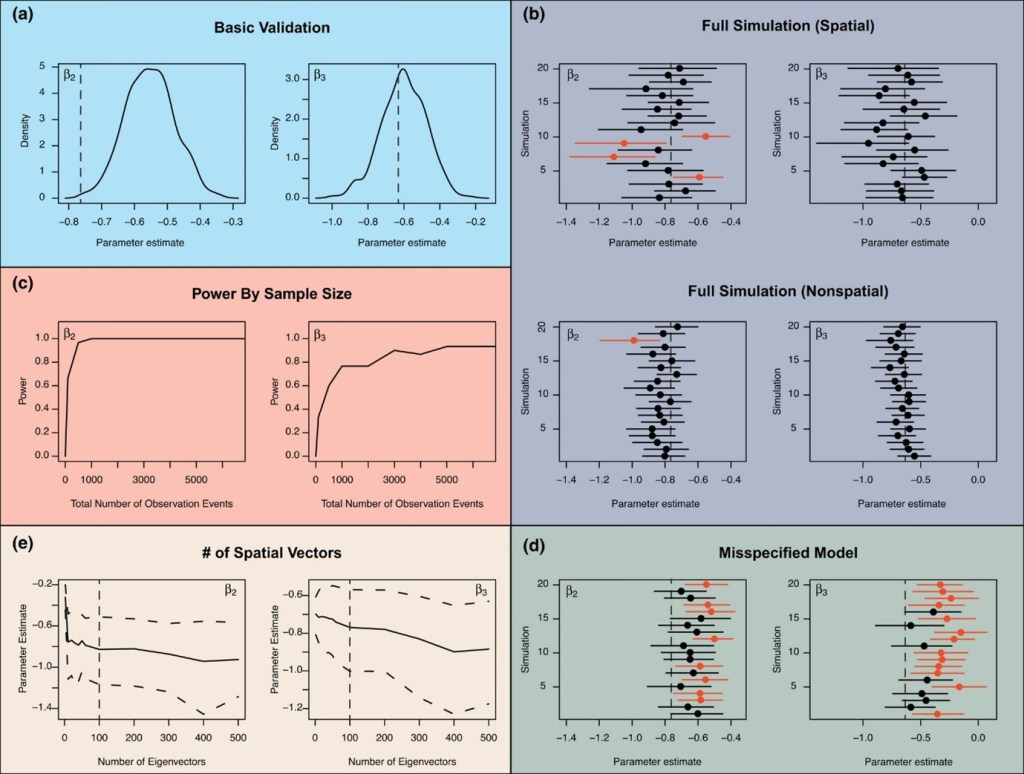
Biologists routinely fit novel and complex statistical models to push the limits of our understanding. software and programs afford the user greater control and flexibility in tailoring complex hierarchical models. However, this level of control and flexibility places a higher degree of responsibility on the user to evaluate the robustness of their statistical inference. This article provides a guide to understanding and validating complex models using data simulations. To determine how often biologists use data simulation techniques
State-space models are an increasingly common and important tool in the quantitative ecologists’ armoury, particularly for the analysis of time-series data. This is due to both their flexibility and intuitive structure, describing the different individual processes of a complex system, thus simplifying the model specification step. This review provide an overview of general state-space models and the different model-fitting tools available.
The bones on the cover
This month’s cover image shows 3D surface scans of the three uppermost bones in the wing of a tawaki (Fiordland crested penguin, Eudyptes pachyrhynchus). 3D surface scans are pre-processed into a lattice of evenly-spaced pseudo-landmark points as an initial step for shape analysis, but with a difference. Here the partial wing is being prepared to be analysed as a single object in a new analytical workflow provided by the R package Morphoblocks. This new R package described by Thomas et al. in this issue provides a workflow for analysing shape variation across whole or partial skeletons. A brief Vignette describing shape change in the upper wing skeleton of penguins is used to introduce the package. This article from Thomas et al. features in the Methods in Ecology and Evolution special feature Realising the Promise of Large Data and Complex Models. ©Daniel Thomas



One thought on “Our January issue is now online and open access!”