BIEN 4.2: A Reproducible Standard for Global Plant Biodiversity Data
Post provided by the BIEN Working Group
For hundreds of years, biologists have carefully collected information on plants, animals, and other organisms and have created and maintained enormous libraries of physical specimens from all around the globe. Specimens are collected with all kinds of information– often there’s a physical example, but beyond that, scientists record where and when the sample was collected, by whom, and what the species name of that organism was at that point in history. Collections of these specimens in museums, herbaria, and other libraries have been essential to scientists studying biodiversity.
But specimens are only one stream of biodiversity data. Over the past several decades, ecological plot inventories, forest censuses, vegetation surveys, and trait measurements have expanded rapidly. Field teams climb into tropical canopies to measure leaf chemistry and branch architecture. Researchers establish permanent plots to track demographic change. Trait campaigns quantify leaf area, wood density, seed mass, and physiological performance across gradients. More recently, citizen science observations have added a new dimension. Together, these complementary but different streams now generate enormous amounts of biodiversity observations across space and time

Big biodiversity data are essential to many grand challenges of the Anthropocene and comparative biology- Historically, specimens may be shipped around the world for direct study. More recently, much of the data associated with physical specimens has been captured on spreadsheets and compiled into large databases. Now, ecologists can download millions of species observations and merge them with trait data, survey data, environmental layers, or genomic resources to map richness, build species distribution models, quantify biodiversity change, and test trait–environment relationships. Biodiversity data in virtual form are now central to comparative biology, modern ecology, and global change biology. These data make analyses possible that were unimaginable even a decade ago.

Nonetheless, the very scale, diversity, biases, and various ways biodiversity data are collected now introduce new scientific challenges and limitations – Geographic metadata must be validated. Outdated or synonymous species names must be reconciled. Provenance must be clarified. Anyone who has tried to use biodiversity data at scale knows that it rarely arrives in a form ready for science. Species names either do not match or are suspect. Taxonomies disagree. Coordinates fall in the ocean or land in curious places. Political divisions are misspelled or inconsistent. Records are duplicated. Some observations come from botanical gardens rather than wild populations. Others are assigned to the geographic centroids of countries or provinces, which makes them look precise when they are not.
A dataset that seems enormous and promising can quickly become a puzzle box of hidden errors, biases, and uncertainty. These frustrations are not rare. They are a routine and growing part of analyzing large biodiversity data.
Big Bad Data? Perhaps Ecologists should not be surprised[1]. In large-scale business analytics, businesses spend around 80% of their time on preparing and managing data for analysis[2]…Similarly, in data science, it’s often quoted that most of the work happens before any algorithm is applied, such that “Data Science is 99% preparation”
The difference is not that biodiversity science is characterized by messy data. It is that we have lacked shared, reproducible standards for resolving these issues. The absence of shared standards for data integration and validation imposes two fundamental constraints on biodiversity science. First, conclusions drawn from large datasets can be biased, and effect sizes can be diluted by unrecognized error. These data inconsistencies propagate through models and forecasts, often invisibly. Second, our science becomes difficult to reproduce because data-cleaning decisions are numerous, rarely standardized, and often not transparent. The concern is that it becomes difficult to determine whether differences across studies reflect biological processes or divergent data sources and preprocessing choices.
If these data issues are not caught early, they can shape scientific conclusions in ways that are hard to detect. Even when researchers are careful, it is often unclear whether two studies differ because of ecology or because of data cleaning decisions that were never fully documented. For example, species richness estimates may be inflated because synonyms are treated as distinct taxa. Range maps can become distorted because a handful of bad coordinates push species into the wrong biomes. Models may become unstable because “native” distributions quietly include cultivated records.
Why BIEN? We built BIEN (the Botanical Information and Ecology Network, see http://www.biendata.org) to avoid having to solve these same data problems over and over again (Figure 2). The result is a suite of analyses and filters that help set a reproducible standard for the biodiversity science community for integrating and cleaning data.
BIEN began in 2008 as a community-driven effort to integrate global data on plant biodiversity. Over time, BIEN became something more than a database. It became an international effort to develop a reproducible workflow ecosystem that makes biodiversity data interoperable, transparent, and reusable.
In our newly accepted paper in Methods in Ecology and Evolution (BIEN: A biodiversity informatics ecosystem advancing open and reproducible workflows for plant observation, plot, and trait data) we describe major updates to BIEN: version 4.2 of the BIEN database, along with an expanding set of tools designed to resolve the most common problems in biodiversity data.
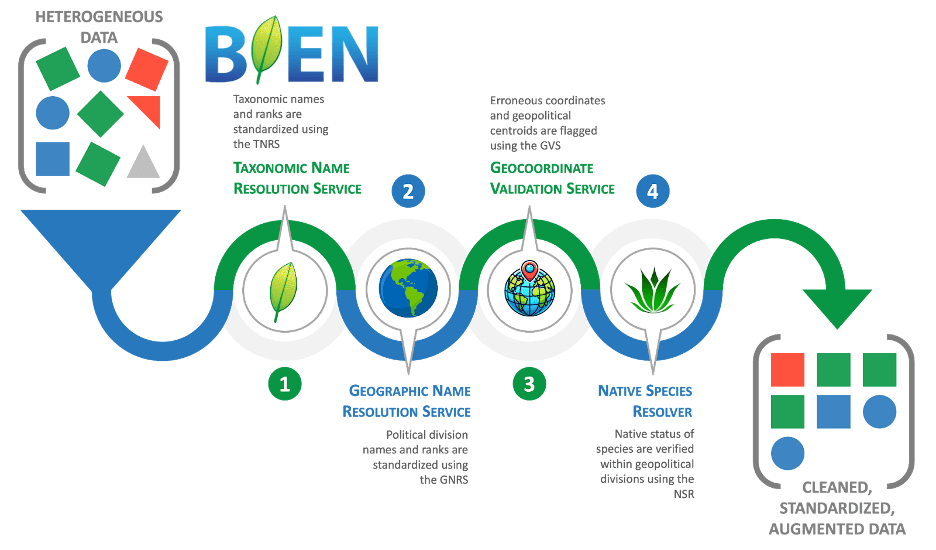
BIEN includes four core data standardization and cleaning services to integrate disparate data sources. Many of these tools can be used independently or together as a pipeline for plant and animal data, as well as other observational data, and are therefore useful for other applications and not specialized to plant occurrences. The end result is that these services can produce analysis-ready datasets.
● The Taxonomic Name Resolution Service (TNRS) standardizes scientific names and resolves misspellings and synonyms. https://tnrs.biendata.org/
● The Geographic Name Resolution Service (GNRS) standardizes political division names across countries and regions. https://gnrs.biendata.org/
● The Geocoordinate Validation Service (GVS) flags invalid or suspicious coordinates, including common centroid errors. https://gvs.biendata.org/
● The Native Species Resolver (NSR) helps distinguish native records from introduced or cultivated occurrences. https://nsr.biendata.org/
BIEN also includes a standardized integrated global database of plant observations that have been passed through the BIEN data standardization pipeline. BIEN 4.2 is now one of the largest standardized plant biodiversity resources available. It integrates:
- 284 million plant occurrence records
- 363,258 ecological plots
- 25,932,454 trait observations
- 54 standardized plant traits
- range maps for nearly 113,000 plant species
These data include herbarium specimens, plot inventories, citizen science observations, and trait datasets assembled from hundreds of studies. These data are now accessible via the BIEN R package. However, BIEN’s most important contribution is not simply scale. It is accessible, quality-controlled infrastructure for plant biodiversity science.
BIEN attaches quality-control information directly to records, allowing users to filter data explicitly rather than relying on hidden assumptions. By doing so, BIEN moves plant biodiversity from fragmented datasets toward reproducible, inference-ready infrastructure that can support conservation planning, macroecology, and global change research.
One sobering lesson from BIEN is how often these checks matter. Even in the largest biodiversity datasets, around 50% of observation records are erroneous in some way – Errors include taxonomic inconsistencies, geographic problems, or uncertainty about whether occurrences represent wild populations. In BIEN 4.2, only about half of the original records pass strict combined taxonomic and geographic validation filters.
Practical uses – A practical example comes from conservation. In collaboration with UNESCO, we used BIEN to estimate the extent of plant diversity within UNESCO World Heritage Sites. This required harmonizing names, validating coordinates, and filtering introduced and cultivated records. Using a fully reproducible BIEN workflow, we estimate that UNESCO World Heritage Sites contain approximately 74,295 plant species, including more than 2,000 threatened species. Without standardization, richness estimates would have been substantially inflated by synonymy, inconsistent taxonomy, and erroneous geographic data. This result highlights why biodiversity informatics is not just a technical detail: it shapes ecological inference and conservation decisions.
In building BIEN, we aim to save researchers time, prevent avoidable errors, and provide tools for reproducible biodiversity science. Despite their foundational role in ecosystems, plants are often underrepresented in global conservation prioritization –frequently reduced to carbon stocks or broad biome categories rather than treated as diverse evolutionary lineages with distinct ecological roles. BIEN helps address this gap by providing standardized, validated plant occurrence, plot, and trait data at unprecedented scale.
We also hope that BIEN can help the ecological community build a shared, reproducible foundation for biodiversity science. Finally, we anticipate that making these data freely available, together with open-source tools, will promote widespread uptake and foster ongoing contributions and refinement from the research community, practitioners, and other interested stakeholders.
Read the full article here.
[1] https://dynamicecology.wordpress.com/2016/08/22/ten-commandments-for-good-data-management/
[2]https://www.forbes.com/sites/gilpress/2016/03/23/data-preparation-most-time-consuming-least-enjoyable-data-science-task-survey-saysdata-cleaning decisions are nuecological factors or because of data-cleaningmerous, rarely standardized, and often /
