We are pleased to announce our Machine Learning Virtual Issue is now online.

This collection of MEE articles showcases exciting advances and applications of machine learning (ML) across a wide range of ecological and evolutionary disciplines.
From the analysis of reef structure and tree crowns, to species and individual animal identification, biological overlap, content analysis, biodiversity assessment and counting animals, ML automates the extraction of meaningful information from large digital collections.
Our Associate Editors Arthur Porto, Marta Vidal-Garcia, Miguel Acevedo, Theoni Photopoulou and Sarab Sethi curated this virtual issue by selecting their favourite MEE articles that use machine learning. Find out below why these papers were chosen, and how they are helping to progress research in ecology and evolution.
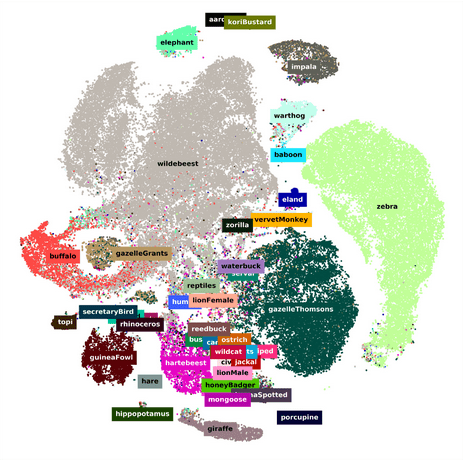
1. Machine learning to classify animal species in camera trap images: Applications in ecology by Tabak et al. (2018)
Miguel: With camera traps the real challenge is that, in theory, somebody has to go, look at them and identify the species. Here the authors developed a highly accurate machine learning model that aids in this process. These machine learning methods coupled with models that account for imperfect detection such as occupancy models are the promising future of wildlife ecology and conservation.
Marta: I chose this exciting application because the authors have developed an R package that allows for ML-based classification of wildlife in camera trap images. I think this package is a great tool for accurately determining the presence or absence of animals in each picture and identifying their species, reducing this time-intensive task and making camera trapping accessible to more ecologists. The authors used an impressive dataset (>3M images) to train their neural network, and found a very high percentage of accuracy during classification, even when using images from other locations.
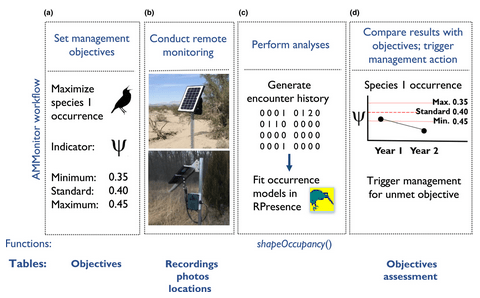
2. AMMonitor: Remote monitoring of biodiversity in an adaptive framework with r by Balantic & Donovan (2020)
Theoni: This paper is interesting because it presents a software framework for moving from data collected by autonomous monitoring units, like camera traps or acoustic recording equipment, through processing, to analysis and interpretation of results. This is a common workflow required by monitoring programs and can be implemented through the R package AMMonitor.
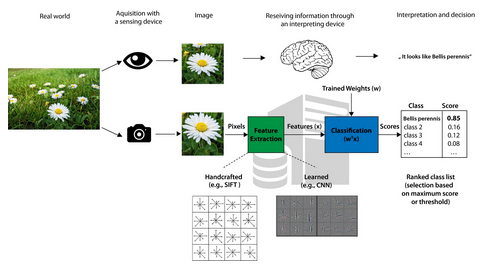
3. Machine learning for image based species identification by Wäldchen & Mäder (2018)
Theoni: In this review, the authors provide a roadmap for researchers interested in applying deep learning neural networks to species identification problems. It helps understand the jargon and presents graphical illustrations of how machine learning works. Although this “what’s what” guide is now two years old, it is still a brilliant starting point.
4. A deep active learning system for species identification and counting in camera trap images by Norouzzadeh et al. (2020)
Theoni: One of the problems with analysing wildlife images from camera traps surveys for biodiversity assessments is that machine learning algorithms learn to recognise the background in images, rather than the subject of interest. Norouzzadeh et al. propose an approach that systematically removes background pixels, increasing correct classification of animals in images, and reducing the need for a human to label a large number of images for the ML algorithm to learn from. The authors also present a clear description of deep learning, artificial neural networks and their structure, and explain the ideas behind transfer learning and active learning.
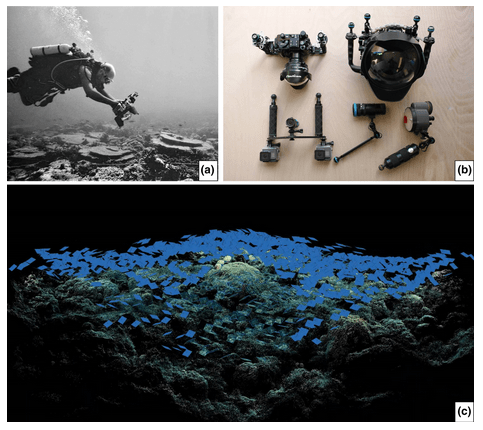
5. A protocol for the large‐scale analysis of reefs using Structure from Motion photogrammetry by Bayley and Mogg (2020)
Arthur: Bayley and Mogg (2020) provide an excellent example of the use of computer vision for the creation of large-scale 3D models of reefs. Given the economic and ecosystem-level importance of these marine systems, this approach has the potential to greatly increase the types of questions that can be quantitatively interrogated and is likely to become a standard tool in ecological surveys.
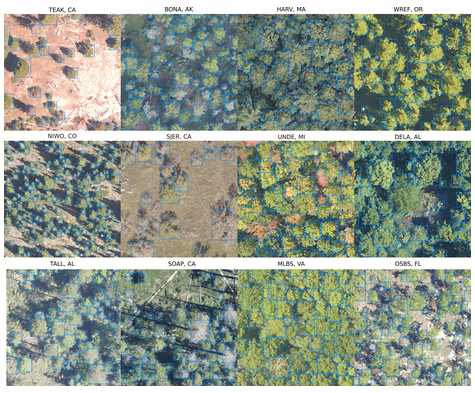
6. DeepForest: A Python package for RGB deep learning tree crown delineation by Weinstein et al. (2020)
Arthur: Weinstein et al. (2020) introduced a new python package for tree crown delineation using RGB images. This is an excellent example of the use of combined hand-labelled and algorithmically generated training data for the training of deep learning models. It also has the added benefit of tackling important questions related to the management of forested landscape. Finally, it is a great example of open science, in which source code, data, and in-development tools are all easily accessible from stable internet archives.
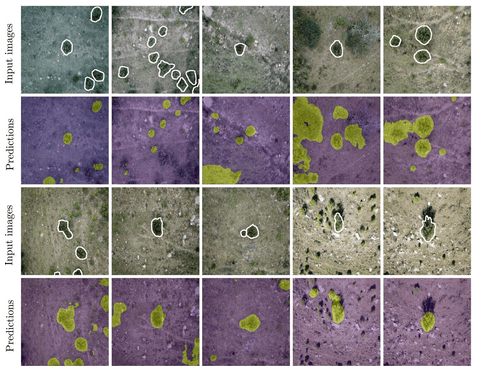
7. Detecting plant species in the field with deep learning and drone technology by James & Bradshaw (2020)
Theoni: An exciting aspect of collecting images through time is that these can be used to detect unusual objects, events or change in ecological features. James and Bradshaw used machine learning to process images from an aerial drone in real time, to identify invasive alien vegetation in South Africa’s fynbos biome. Invasive woody plant species are a problem in the fynbos biome because they outcompete the indigenous vegetation, resulting in dense stands of alien plants. This prevents recovery of indigenous species through crowding and exacerbates the effects of drought. Detecting the spread of alien vegetation early can make a real difference to how easy it is to control.
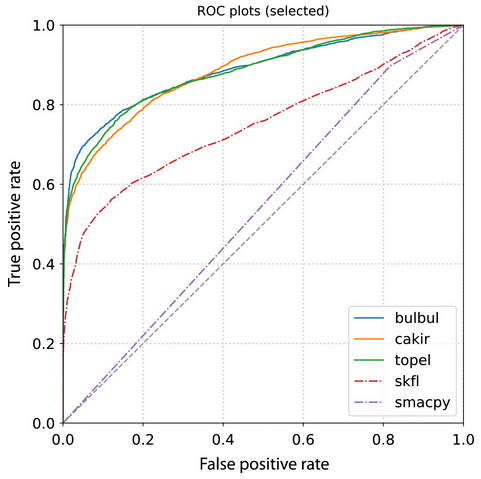
8. Automatic acoustic detection of birds through deep learning: The first Bird Audio Detection challenge by Stowell et al. (2018)
Miguel: Some argue that acoustic sensors and camera traps are the future of wildlife ecology. These are transforming the way we study wildlife ecology at large-scales. In this paper, the authors developed a “Bird Audio Detection challenge” where they “challenged” different groups to come up with the best machine learning method to identify bird species from recordings. Out of these, many performed well with high accuracy.
Sarab: Acoustic species monitoring is, perhaps, uniquely able to excite and terrify ecologists at once. The potential for continuous, real-time species inventories will only possible once reliable species detection models are widespread. The authors of this work collated a diverse array of labelled datasets, and challenged the ML community to design robust detectors that worked across a wide array of situations. The outputs from this challenge has set the bar for the field, and the winning model, ‘bulbul’, is still used widely three years on (a lifetime in the ML world!)
9. Deep learning‐based methods for individual recognition in small birds by Ferreira et al. (2020)
Arthur: Ferreira et al. (2020) provides an interesting computer vision approach applied to individual identification of birds, both in the lab and in the wild. This study’s key contribution is the development of an analytical pipeline that allows training of deep learning models for individual recognition without the need for external markers. With this pipeline in hand, monitoring of free-range biological populations becomes a much more accessible goal.
10. Wavelet filters for automated recognition of birdsong in long-time field recordings by Priyadarshani et al. (2020)
Theoni: This paper presents a comparison of a wavelet approach and machine learning approaches applied to the classification of birdsong for species identification. I like this paper because of the methods comparison and the fact that the authors’ chosen approach can either be used in place of a machine learning approach or, generate input for a machine learning approach. This highlights the need to think carefully when choosing an approach for a classification task.
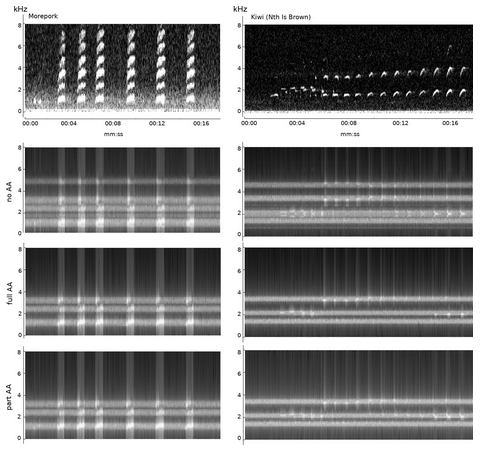
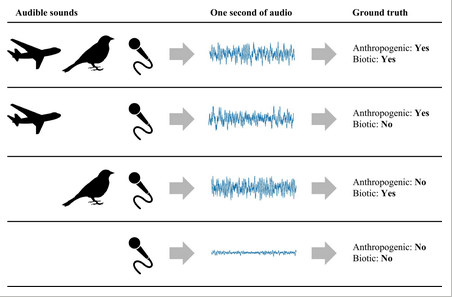
11. CityNet—Deep learning tools for urban ecoacoustic assessment by Fairbrass et al. (2018)
Theoni: Although the focus has been on detecting and identifying natural sounds, produced by animals, there is a growing need to understand the effects of human-made noise on wildlife. Before we can understand the effect to human-made noise, we need to be able measure it, as well as measure natural sounds, sometimes at the same time. In this paper, the authors develop an approach for measuring both audible biotic and anthropogenic acoustic activity in urban environments for an acoustic assessment of biodiversity in cities.
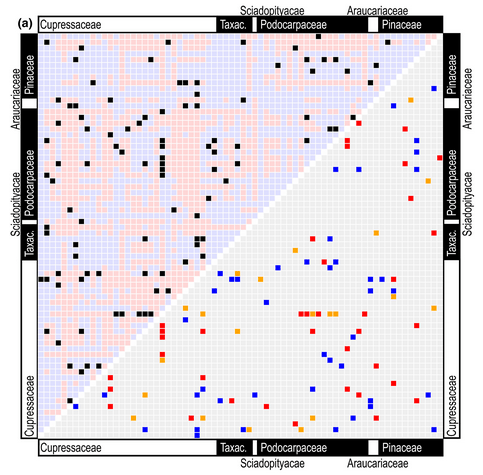
12. hyperoverlap: Detecting bio- logical overlap in n-dimensional space by Brown et al. (2020)
Theoni: There is a strong focus on detecting similarities in sounds and images so that we can match or label records as the same species or the same individual. However, there are some cases where failure to detect similarity, or mistakenly detecting similarity, can be informative. Brown et al. used misclassification of morphological or ecological data to detect overlap between species or populations in comparative biological studies. Their approach used Support Vector Machines (SVMs) to first train a classifier on labelled data from two groups and then find the optimal boundary between them. They demonstrate the approach for comparing the climatic distribution of conifer genera, reducing the computation time by 80%. A real advantage of this approach is that it is especially effective for large numbers of comparisons even when the numbers of occurrences within individual groups is small.
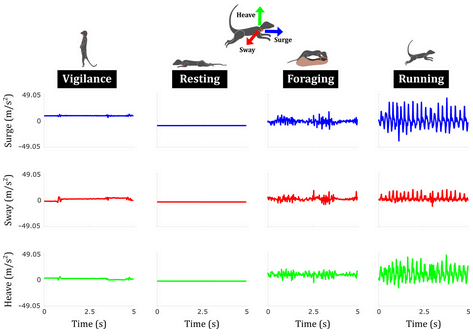
13. Seek and learn: Automated identification of microevents in animal behaviour using envelopes of acceleration data and machine learning by Chakravarty et al. (2019)
Marta: The authors used an ML approach, along with their knowledge on biomechanical principles that characterise movements, in order to classify animal behaviours using accelerometers. The exciting part about this approach is its potential for allowing classification of behaviours in less ideal study systems. For example, it could be very useful for secretive or endangered species, in which it could be used to detect changes in behaviour without an observer effect (possibly affecting the data), and to detect personality traits, based on combinations and amounts of certain behaviours.
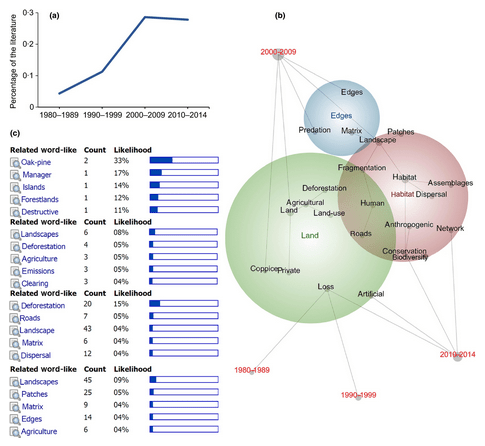
14. Automated content analysis: addressing the big literature challenge in ecology and evolution by Nunez-Mir et al. (2016).
Sarab: Most scientists can attest to the difficulty of sifting through huge swathes of prior art in any given field. In this beautifully illustrated meta(-meta?)-study, the authors introduced automated content analysis (ACA) as a tool that can allow ecologists and evolutionary biologists to summarise and structure enormous volumes of literature in an efficient manner. Synthesis on this scale can offer unrivalled insight into how fields intertwine, potentially even revealing new exciting directions for future work.
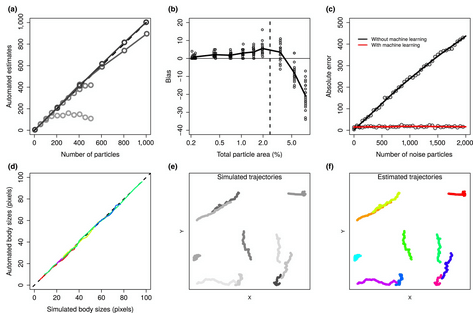
15. trackdem: Automated particle tracking to obtain population counts and size distributions from videos in r by Bruijning et al. (2018)
Theoni: Population assessment is one of the fundamental yet non-trivial cornerstone issues in ecology. Most of ecology either hinges on or is trying to get at how many animals are in a population, and what their energetic status is. In this paper, Bruijning et al. present a method that can extract individual size information and population size from video data – one of the most human-time-intensive information types. Combined with individual identification, where this is possible, this approach has the potential to be a powerful ecological tool that could feed into classical statistical methodology for wildlife population assessment.






great explanation…thank you for sharing with us
Nice Post…. Thanks for sharing this great information with us.
dimensionality reduction techniques
what is dimensionality reduction