Robert May Prize Shortlisted Article
Post provided by Alberto Pascual-García
Each year Methods in Ecology and Evolution awards the Robert May Prize to the best paper in the journal by an author at the start of their career. Alberto Pascual has been shortlisted for his article ‘functionInk: An efficient method to detect functional groups in multidimensional networks reveals the hidden structure of ecological communities’. In this post, Alberto discusses the application of the functionInk (functional linkage) package for distinguishing modules and guilds from large multidimensional networks.
Biology and dialectics, two faces of the same coin
We need concepts to grasp the reality around us. But, as Hegel noted, “natura non facit saltum” (there are no leaps in nature) which means that in Biology boundaries are difficult to delineate and so, when it comes to studying biological systems, conceptualization becomes a major challenge.
Having a background in physics, when I started my Ph.D. in biology I soon realized that biological systems require a closer look into our epistemological beliefs. Many frequently used concepts, such as emergence, are often poorly defined. Georgescu-Roegen noted that there are concepts that can mean one thing and its opposite at the same time. Since these concepts apparently violate the Law of non-contradiction, a cornerstone in Hegelian philosophy, he suggested calling them dialectic.
Chief among dialectic concepts in biology is the notion of “function”, because it is difficult to define it without using a finalistic explanation. Defining what function is may require talking about its opposite (when or how function ceases to be), and a fundamental reason behind the dialectic nature of biological concepts is that biological systems are made by components entailed by interdependent relations determining “what they do and to what end”.
In this respect, the Rashevsky-Rosen school on relational biology encourages us to investigate biological systems “Throwing away the matter and keeping the underlying organization.” In other words, the details about the elements in a biological system are secondary to their relationships. This is possibly why complex networks theory has become a very popular mathematical framework to investigate biological systems during this century.
How relations speak about dialectic concepts
During my Ph.D., I worked on very different biological questions and I used complex networks in most of them. I learned that one of the most powerful features of complex networks is that it provides an empty-of-content framework which is so flexible that we can study how sets of elements of any kind are connected through any kind of relationships. But perhaps the most exciting feature of the framework is that, when certain properties are analysed, one can deal with (often dialectic) concepts having different meanings depending on the data.
Consider a first example in which we have three genes: A, B, C. We set a threshold by which, if any pair of genes have a sequence identity of 70%, we can argue they have a shared ancestry. This threshold is of course somewhat arbitrary: it doesn’t mean that a pair of genes can’t share ancestry if their sequence identity is 69.9%. There is some vagueness in our definition. Despite this, if we build a network and we find that the pairs A ~ B, and B ~ C have a similarity well above 70%, then we can infer that A and C also have a shared ancestry, even if their similarity is well below than 70%. How? The network is showing us the pathway: if the genes are related through duplication and subsequent divergence, then homology is transitive (i.e. a relative of one genes’ relative is its relative). And voilà, we circumvented the vagueness in the similarity threshold.
The property we used in the example is ‘transitivity’: if A ~ B and B ~ C then A ~ C (“if A is related with B and B is related with C, then A is related with C”). And it is a property implicit in several complex networks metrics like the clustering coefficient. In a second example, consider that the elements A, B and C are bacterial species, and the relationship A ~ B indicates that “A outcompetes B” when grown together. If, when analysing the network, we observe that A outcompetes B, B outcompetes C, and C outcompetes A, we can deduce that, under certain conditions, the three species could coexist.
In these examples, homology and coexistence are predicted from the analysis of the networks. They are not encoded in the information provided by each specific link, they are rather complex concepts that “emerge” from the network properties, in these examples from the transitivity property.
Community structure in complex networks
Other network properties may inform us about different complex concepts, depending on the elements and on the relations considered. A central question in complex networks is if there exist community structure, namely a group of nodes sharing some characteristics in common. The most widely used definition of community are “modules”, which are sets of elements (nodes) tightly connected between members of the same community and loosely connected with elements of other communities.
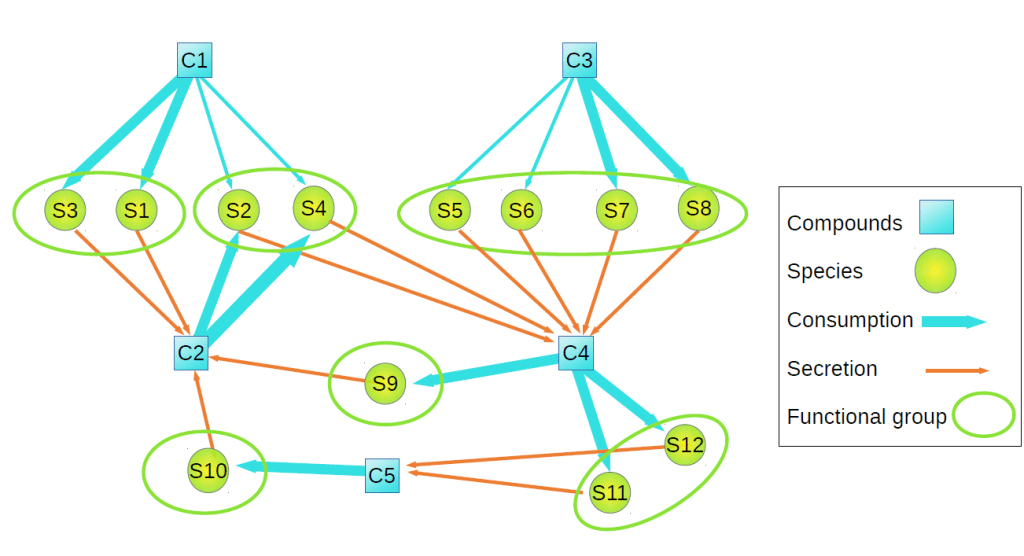
However, when my collaborator Thomas Bell and I were working on networks of bacterial communities, we realised that the concept of ‘module’ was not the most relevant for certain biological questions. To illustrate this point, consider a network connecting bacteria with the resources they consume and secrete, and that we aim at identifying communities of bacteria that consume and secrete roughly the same compounds (see illustration below). Bacterial communities fulfilling this definition are “functionally equivalent” from the point of view of their metabolism, and they form a guild.
The notion of modularity would not help us finding guilds, because bacteria are not directly connected in the network: the network is bipartite, namely we have two types of nodes, bacteria and resources, and nodes of the same type are not connected. So we asked ourselves: Is it possible to find an empty-of-content definition of function for a node in any arbitrary network and identify functional groups? In other words, Is it possible to say what the function of a node is in the context of a network, independently of the meaning of the node (e.g. bacteria or gene) or the nature of the relationships between nodes (e.g. ecological interaction or shared ancestry)? And if so, is it possible to interpret these functional groups as modules or guilds depending on the context?
From functional roles to functional groups
To answer this question we borrowed the concept of “role” in social networks. The influence of an individual in a social network depends on the nature and number of her relationships, and on the specific individuals she interacts with. This definition is flexible enough to accommodate different types of relationships. This is important because, nowadays, we deal with large datasets in which different types of information are considered in a single network. Therefore, the “role” should include the fact that a node may have different types of links, with different meanings. Following this reasoning we realized that, independently of what a node or a link represents, what is clear is that when two nodes have the same number and type of connections with the same neighbours, they play the same functional role in the network –the “functional groups” in the illustration above.
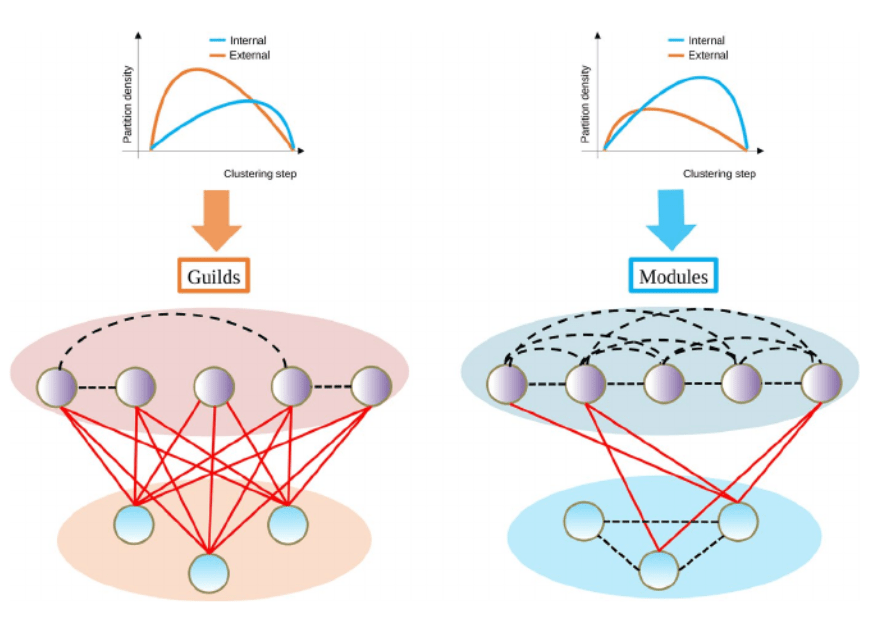
We proposed metrics comparing how similar any pair of nodes are in terms of their connections, from which we can simply cluster nodes leading to communities that can be interpreted as functional groups. The final step of the process is finding at which step of the clustering we find “optimal” communities. And because there is a “No Free Lunch” theorem for optimality in community detection methods (no method is optimal detecting communities in all networks), this was a challenge. Instead of looking for a single “optimal” cut-off, we developed two metrics that allow us to evaluate which kind of communities represent a better dimensionality reduction of the network. In the end, we noted that both types of communities (modules and guilds) respond to the definition of functional group we proposed. Since we are clustering (linking) nodes according to their functional role, we called our method “functionink” (functional linkage).
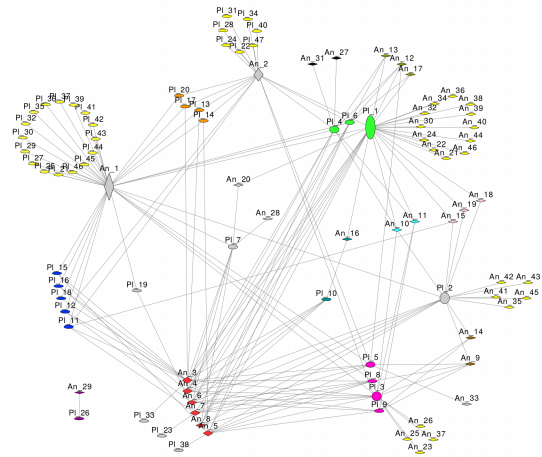
Perhaps the most interesting feature of our method is that the communities found have a straightforward interpretation, because the metric comparing the nodes and the clustering are very intuitive (yes, you don’t need to understand what a Laplacian matrix or a stochastic block model is). For instance, the nestedness in plant-pollinators networks is a network property that sparked tremendous controversy in the literature. On the one hand, because it has been largely discussed if it is a non-random pattern. On the other hand, because it has connections with several dialectic concepts, most notably the stability of ecosystems. One of the most remarkable features of the pattern is that specialist species tend to be connected with generalists species. But it is possible to gather a much better intuition of this metric by identifying functional groups (see illustration below). Beyond the clear connection between specialists and generalists, we immediately observe other interesting features, such as a core of three highly connected functional groups. We can also speak about functional redundancy of species, because species between the same functional group are functionally equivalent. As a corollary, we can foresee if the consequences that the removal of one species in the system will have are more or less dramatic, depending on the functional group it belongs to. In summary, it disentangles functional building blocks of the nestedness pattern, which may help us to make its connections with other concepts more concrete.
Summary
In the article, we present and interpret some examples coming from plant-pollinator, trophic and microbial networks, and we found that it is possible to shed light on elusive concepts related with ecological function, such as the definition of ecotypes. This is why we believe functionink will be a valuable tool to explore other dialectic concepts in ecology, evolution and beyond… hopefully from your data!
Find out more about the articles that were shortlisted for this year’s Robert May Prize here.