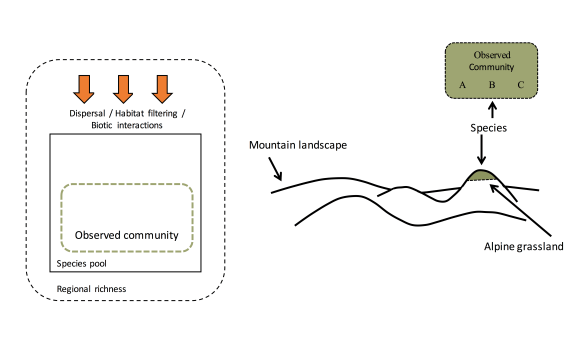
The quest for a sharp definition of function in biological networks
Robert May Prize Shortlisted Article
Post provided by Alberto Pascual-García
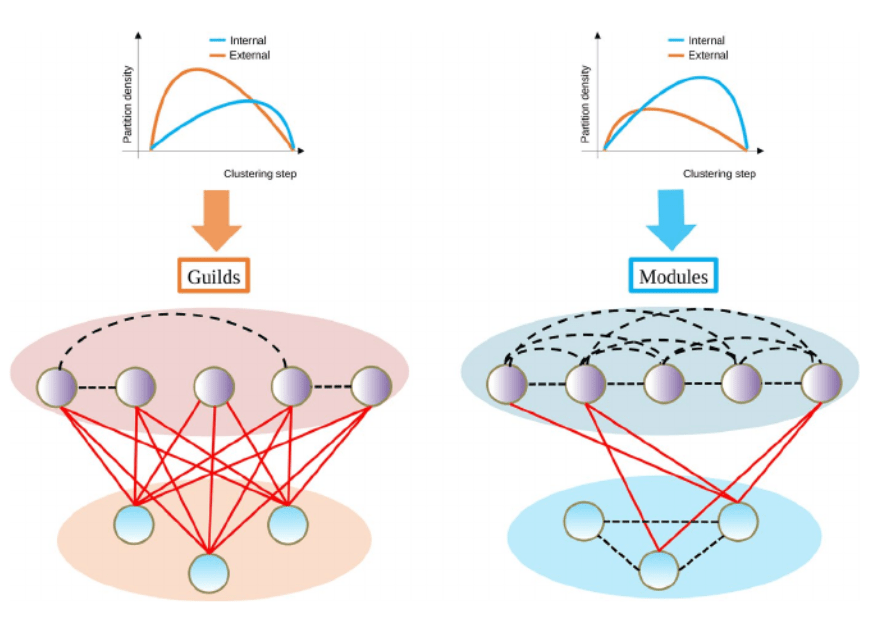
Each year Methods in Ecology and Evolution awards the Robert May Prize to the best paper in the journal by an author at the start of their career. Alberto Pascual has been shortlisted for his article ‘functionInk: An efficient method to detect functional groups in multidimensional networks reveals the hidden structure of ecological communities’. In this post, Alberto discusses the application of the functionInk (functional linkage) package for distinguishing modules and guilds from large multidimensional networks.
Continue reading “The quest for a sharp definition of function in biological networks”