Our May Issue is now online! This issue contains 19 fantastic articles about the latest methods in ecology and evolution, including methods for investigating animal movements, predicting species co-existence patterns, exploring primary production in macroalgal canopies and much more!
Read on to find out about this month’s featured articles and the article behind our beautiful bee and sunflower cover.
Featured Articles
PhycoCanopy *open access* Macroalgal canopies are considered important for coastal food webs and may have a role in carbon sequestration. Until recently, measures of canopy photosynthesis have been relatively rare, and simulations have sometimes omitted key aspects. Here, Mark P Johnson presents the R shiny tool PhycoCanopy, which offers a way of exploring how different algal parameters and environmental settings can affect net canopy photosynthesis.
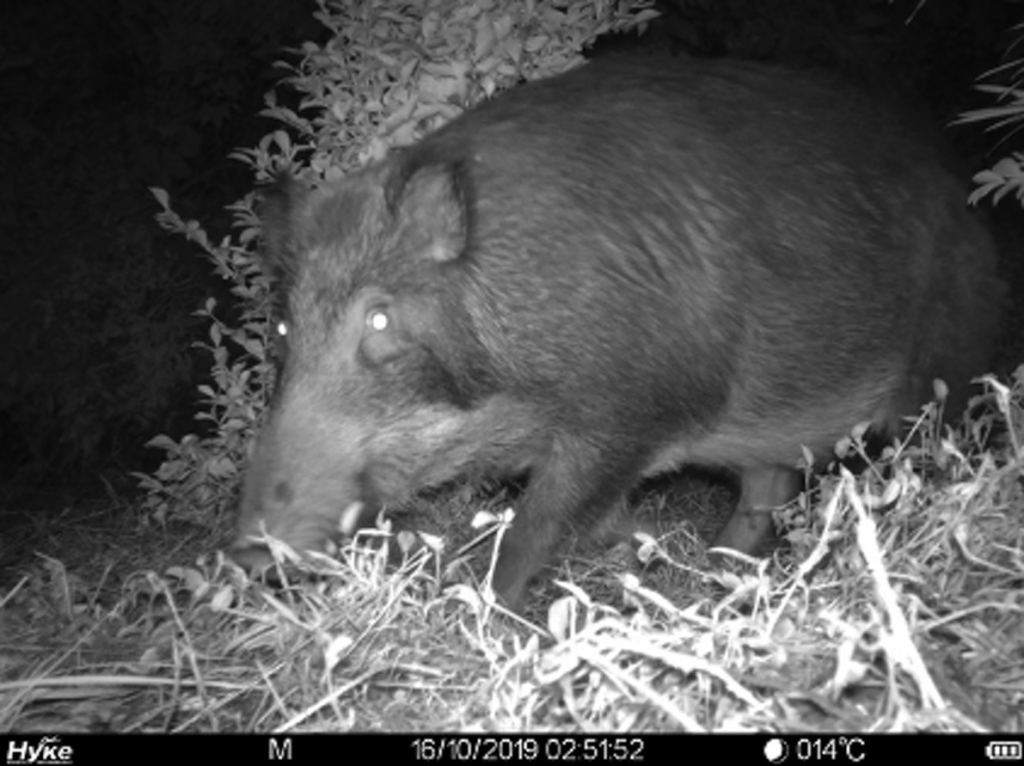
Unmanned aerial vehicles as a useful tool for investigating animal movements *open access* Determining animal abundance is crucial for assessing the effectiveness of management measures against pest animals, and investigating animal movements has become important for conducting abundance estimations of unmarked animals. Here, Iwamoto et al. propose a method for estimating animal movements using unmanned aerial vehicles, conducting a case study that aimed to investigate the movement of wild boars.
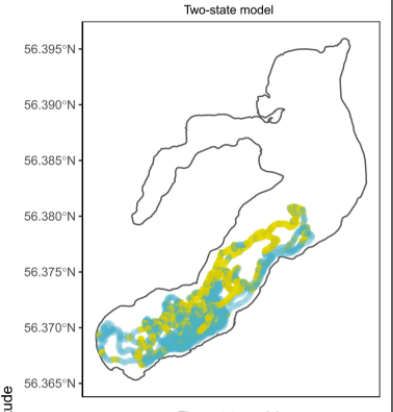
Predicting aquatic animal movements and behavioural states Fine-scale tracking with passive acoustic telemetry can yield great insights into the movement ecology of aquatic animals. To predict fine-scale positions of tagged animals in continuous space from spatially-discrete detection data, state-space modelling through the R package YAPS provides a promising alternative to frequently used positioning algorithms. However, YAPS cannot currently classify multiple kinds of movement that may be used as proxies for individual behaviours of study animals (behavioural states), an endeavour that is of increasing interest to movement ecologists. Here, Whoriskey et al. advance YAPS by incorporating the functionality to predict behavioural states by using an iterative maximization framework.
Predicting species coexistence patterns Predicting coexistence patterns is a current challenge to understand diversity maintenance, especially in rich communities where these patterns’ complexity is magnified through indirect interactions that prevent their approximation with classical experimental approaches. Here, Hirn et al. explore cutting-edge Machine Learning techniques called Generative Artificial Intelligence (GenAI) to predict species coexistence patterns in vegetation patches, training generative adversarial networks and variational AutoEncoders that are then used to unravel some of the mechanisms behind community assemblage.
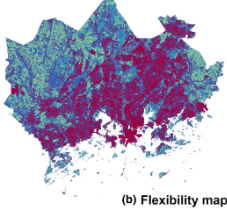
Novel methods for spatial prioritisation Spatial prioritisation integrates data on the distributions of biodiversity, costs and threats. It produces spatial priority maps that can support ecologically well-informed land use planning in general, including applications in environmental impact avoidance outside protected areas. In this article, Moilanen et al. describe novel methods that significantly increase the utility of spatial priority ranking in large analyses and with interactive planning.
The Bee on the Cover
A golden northern bumble bee, Bombus fervidus, forages on Helianthus annuus. Numerous species of bumble bees, important pollinators of agricultural and natural ecosystems, have experienced recent population declines. Improved methods, such as terrestrial eDNA techniques, are needed to monitor the abundance and distributions of such species. In this issue, Richardson presents statistical procedures for controlling critical mistag-associated false discoveries within metagenetic data. This work is particularly relevant to use cases where species presence is inferred from minimal numbers of metagenetic sequences, as in the case of pollinator community surveillance using eDNA methods. In this context, most sequences represent non-target organisms while the species of interest account for a minuscule fraction of sequencing coverage and can be difficult to distinguish from sequencing artifacts such as contaminants and critical mistagging events. Statistical methods for controlling mistag false discoveries improve the robustness of metagenetic techniques and expand their applicability into new research contexts. Photo credit: ©R. Richardson