Post provided by Narimane Chatar (She/Her) Romain Boman (He/Him), Valentin Fallon Gaudichon (He/Him), Jamie A. MacLaren (He/Him), Valentin Fischer (He/Him).
Understanding the way that bones and other biological materials deal with the stresses and strains of everyday life is fundamental for interpreting the behaviour of modern and extinct organisms. Researchers frequently do this by using a digital simulation which can predict the behaviour of materials by breaking complex objects down into much smaller elements – this is known as finite element modelling. In this blog post, Narimane Chatar and her co-authors discuss their new protocol for performing finite element modelling aimed at life-sciences and biomechanics which is fast, open-source, and free for all to use.
Although initially developed for civil engineers, finite element analysis (or FEA) was quickly incorporated into biological and palaeontological sciences to predict the behaviour of structures in extinct and extant organisms. Such structures may range from bones and teeth of vertebrates and exoskeletons of invertebrates to the stems of plants. In animal sciences, predicting the behaviour of structures which are moved by muscles can be tricky, as muscle is an extremely complex structure to model. There are very few protocols available which can realistically simulate the way in which muscle fibres wrap around bones.
In this post, we discuss our new finite element protocol which can perform muscle-induced biomechanical simulations. Using FEA for studying the mechanical behaviour of modern and extinct organisms is particularly useful as it is non-destructive – all the stresses and strains are simulated in a virtual environment with no damage to actual materials (such as one-of-a-kind fossils).
The first step was to digitize the object of interest in three-dimensions. The 3D representation of the structure could be obtained through surface scanning (e.g. using a laser), or by using computed tomography (CT) scanning which can also provide details of the internal geometry of the object. Relevant forces need to be applied onto the structure to simulate the action of muscles, and that is where things normally become much more complicated. Our automated procedure builds the structure, defines important constraints on the 3D model, and distributes muscle forces across the model all within a single, free and open-source simulation software called FOSSILS.
Muscle induced simulation: a more realistic way to simulate a movement
Muscle is a complex structure to model. As a result, biologists and palaeontologists simulating the action of muscles on other materials will often model muscle forces indirectly, by using the assembled forces in positions corresponding to the direction of load (load vectors), or the points between the simplified elements making up the 3D model (focal nodes). However, even when indirect forces are used, the modeling of muscle action on the entire area of insertion remains more accurate.
One of the difficulties of modeling muscle forces on their area of insertion comes from the manner in which muscle fibers wrap around or across a bone. Solving this conundrum was first done in 2007 with a computational routine called BONELOAD, which was developed by Prof. Grosse and colleagues from the University of Massachusetts to distribute muscle forces onto a surface defined by the user.
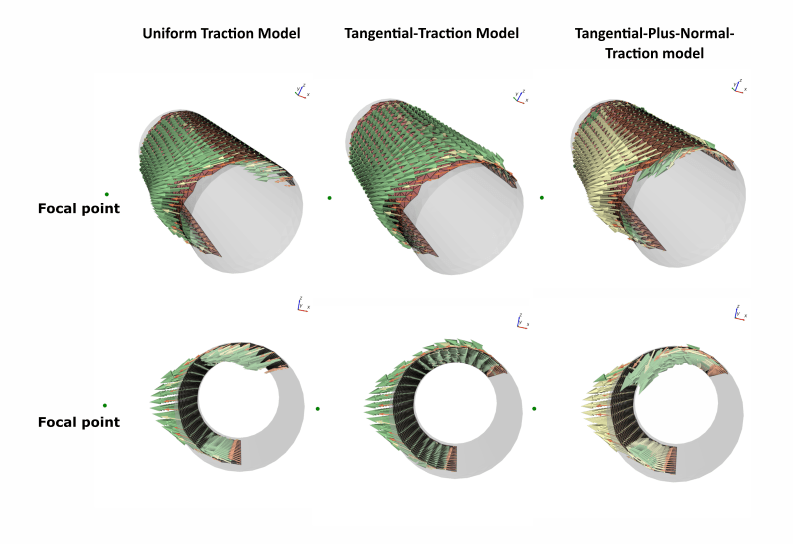
We updated this routine and created an open-source linear static solver, implementing three different ‘traction scenarios’ which distribute forces across a muscle insertion area. The first scenario is called a Uniform Traction. In this scenario, force vectors pass straight through the surface of the bone. This is not entirely realistic, but does reduce computation time for the model. The second scenario is a Tangential Traction, which simulates the wrapping of muscle fibres around the bone without any force vectors passing through the bone – this is much more life-realistic than the Uniform scenario. The last scenario is a Tangential-Plus-Normal Traction scenario. This model not only simulates the forces within the muscle fibres wrapping around the bone, but also incorporates the contact forces between muscle and bone (the normal force) which is also pulling on the bone. The Tangential-Plus-Normal model thus represents the most life-realistic muscle-muscle modelling simulation available.
A wide range of possible applications
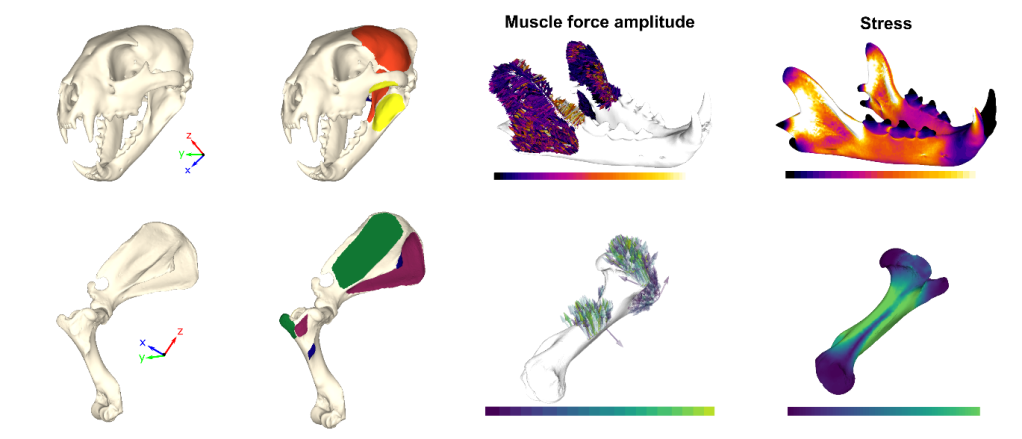
The application of FEA in life sciences is highly variable, and can be used to simulate a wide range of behaviours including biting (for feeding or fighting), locomotion, breathing, etc. To demonstrate the flexibility of our protocol, we used it in three different simulations: two different biting scenarios on jaws from different vertebrates (an extinct marine reptile and a modern leopard) and a shoulder rotation in a modern hooved mammal (a tapir). In addition to running models rapidly and with high accuracy, FOSSILS also provides an additional aesthetic make-over for finite element analyses by offering several colour palettes, some of which are much easier to interpret for colour blind people!

FOSSILS is compatible with multiple 3D file formats and only requires a few processing steps that can be performed in various other 3D processing software (including freeware such as MeshLab, Blender, etc.). FOSSILS writes the resultant force outputs in a very simple format which can easily be extracted and translated into another format for statistical analysis and dissemination.
Comparison with existing solutions
Finite element modelling software are commonplace in physics and engineering institutions or university departments. However, these computer programs often require costly licenses which need renewing each year and very expensive and powerful computers to run the simulations.
During our study we compared FOSSILS to some existing, commercial finite element solutions. We performed the same analysis on our three examples using a commercially available protocol Metafor, the widely used MATLAB/Strand7 software combination, and our new FOSSILS protocol. We found that the results were consistent across the three finite element codes compared. We were also able to demonstrate that FOSSILS represents not only a comparable solver, but a much faster alternative to commercially available alternatives – up to four times faster! We hope FOSSILS will enable muscle-driven finite element analyses to be cheap, swift, and accessible to a wide range of researchers from diverse scientific, methodological, and financial backgrounds.
One-step further: Muscle induced simulations on multi region mesh
Our protocol in its current form has the possibilities for modelling individual objects (i.e. single bones with uniform properties) in a biomechanical simulation. A few years ago, the ‘GenIso Mesher’ – a method for simulating multiple regions within a mesh – was implemented in Metafor, the parent software of FOSSILS. This enabled the generation of a 3D surface mesh from a CT scan (for example using the 3D Slicer software.
More specifically, GenIso allows the user to apply different mechanical and behavioural properties to different parts of the 3D model. As a result, GenIso can be used to model bone with different internal (trabecular) and outer (cortical) regions and corresponding material properties which simulate their behaviour under loading in real life scenarios. GenIso was initially implemented in Metafor, however, it can be coupled with our FOSSILS protocol to provide an even more life-realistic finite element solver which remains fast and affordable.

The source code and installation files for FOSSILS, and all the input files for running the same simulations as were produced in our recent study, are openly available in the supplementary material for the paper (mee314051-sup-0001-Supinfo.pdf), as well as on the GitLab server of the University of Liège and on Zenodo. Readers can find more details on GenIso at http://hdl.handle.net/2268/136159.
You can read our full Methods in Ecology and Evolution article by following the link below: