post provided by: gustavo burin
Este post também pode ser lido em Português
A couple of days ago I came across a nice video (in Portuguese only, sorry) about so-called “living fossils”. The video focused on the problems of using them as arguments against evolution. But I’d like to take the opportunity to talk more about these long-lived lineages.
‘Living fossil’ is a term used to describe lineages that are thought to have been around for a very long time and retain characteristics that resemble of their fossil relatives. A couple of well-known examples of these lineages are the Tuatara of New Zealand (Sphenodon punctatus) and the Gingko tree (Gingko biloba).
How can we ‘Identify’ a Living Fossil?
Usually, these lineages show very long branches in molecular phylogenies, much longer than current lineages typically show (see the example below). The length of these branches are proportional to time (usually in millions of years), so the longer the branch the longer the lineage has lived. These very long branches could give us the impression that certain species are present for even hundreds of millions of years.
But, in reality, they’re are probably much, much younger than that. These very long branches are most likely artifacts from the way these phylogenies are built. To better understand this, we need to take a step back and review how a molecular phylogeny is made.
Molecular phylogenies, for the most part, are made using only DNA data from extant species (since it is almost impossible to obtain reliable DNA information from fossils). As in any other study in ecology and evolution, to build an informative and reliable phylogeny we need to have information from as many species as possible. Otherwise, both the relationships between species and the tempo of their divergences could be incorrectly reconstructed and everything that follows would also be incorrect. So, if the number of species with available molecular data is broad enough (corresponding to a significant fraction of the total richness of the group), molecular evolution models can help us to understand how DNA sequences of all those species have evolved from the most recent common ancestor (MRCA) of the entire group.
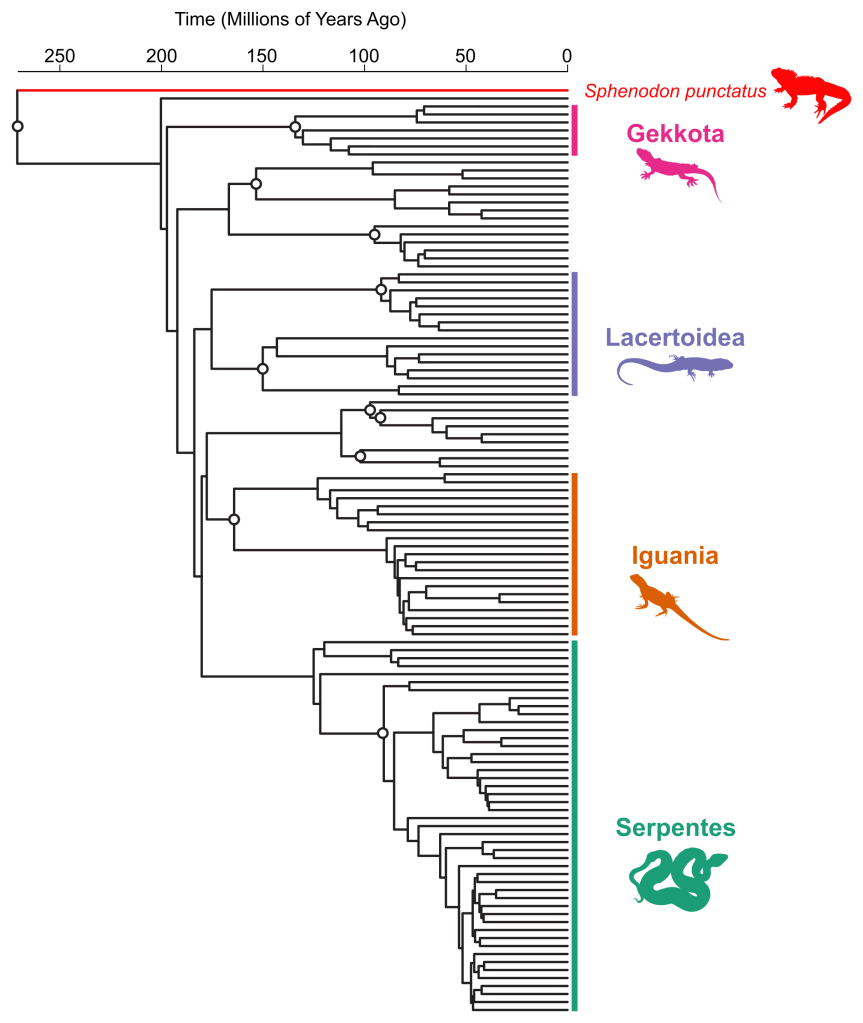
Living fossils belong to lineages that diverged a long time ago from its other ‘non-living fossils’ relatives. Due to this ancient divergence, we’d expect that their DNA would reflect this evolutionary history and that those living fossil lineages would be attached to the base of the phylogeny. That would mean those lineages are represented as originating a very long time ago, with very long branches (up to a couple hundred million years old in the case of the Tuatara, for example). But, it’s important to keep in mind a key limitation of molecular phylogenies: they don’t record direct information about lineage extinction!
As I mentioned before, they’re built using information from extant species only (although fossil information can be used for time calibration). This means that lineages that have originated and gone extinct won’t be represented in the resulting phylogeny at all. This absence gives the impression that nothing happened along all the time the living fossil ‘existed’.
Are Living Fossils ‘Highlander species’?
We know from the fossil record that the average duration of mammal species, for example, is between 2 and 4 million years (although this varies quite a lot between different groups but let’s use this value as a ballpark). With that in mind, we’re practically forced to ask “what on earth do those living fossils have that allow them to live 20x longer than a typical species? Are they the missing link that connects typical vertebrates to the MacLeod lineage?” The answer to both of these questions may lie in extinction.
Let me explain: we know that long branches in molecular phylogenies can be generated through long histories that consisted of several extinction events. These events erase information on species that originated and went extinct throughout the evolutionary history of the group.
Let’s use some simulations of macroevolutionary processes to illustrate this: both phylogenies shown below are representations of the same simulation . The phylogeny on the left contains all species (including the extinct ones). The one on the right contains only the extant species. We can see that some of the branches look much longer on the phylogeny on the right.

When looking at those long branches, our intuition makes us think (at least at first) that nothing (no speciation or extinction) happened along them. Based on our understanding of the evolutionary dynamics though, the better interpretation is that a lot of things might have happened. Many different species were probably generated in such a long time period. Most of them went extinct and only one (or very few) are left alive today to tell the story of the entire lineage. Due to this story of speciation and extinction, the species that are still alive today most likely didn’t originate many millions of years ago.
This is one more reason why the term ‘living fossil’ isn’t accurate. Besides the diversification dynamics, another possible explanation for the similarity between the extant species and their fossil relatives can be found in low rates of morphological evolution. But that’s a subject for another time.
If you want to talk more about macroevolution, hit me up on twitter!
To find out more, read our Methods in Ecology and Evolution Virtual Issues ‘Advances in Phylogenetic Methods’ and ‘Phylogenetics and Comparative Methods’.



One thought on “The Problem with ‘Living Fossils’: A Molecular Phylogenetic Perspective”