Our February issue is now online! Our second issue of the year contains 22 high-quality articles about the latest methods in ecology and evolution. This month we have methods for visualising the tree of life, estimating arthropod abundance and diversity, disentangling effects of climate and land use on biodiversity and much more! This issue also contains four Applications and two Practical Tools articles that are free to read for the next two months. Read on to find out about this month’s featured articles.
Featured Articles
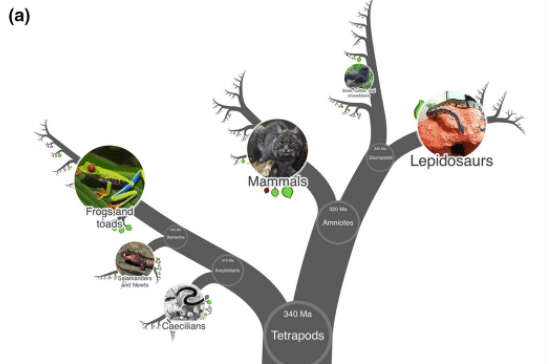
The OneZoom project *open access* The complete tree of life is now available, but methods to visualise it are still needed to meet needs in research, teaching and science communication. For the OneZoom project, Wong & Rosindell automated data processing, synthesised data from a wide range of available sources, then fed these data to a client-side visualisation engine in parts. They developed a way to store the whole tree topology locally in a highly compressed form, then dynamically populate metadata such as text and images as the user explores. The result is a seamless way to explore the complete tree of life, including images and metadata, even on relatively old mobile devices.
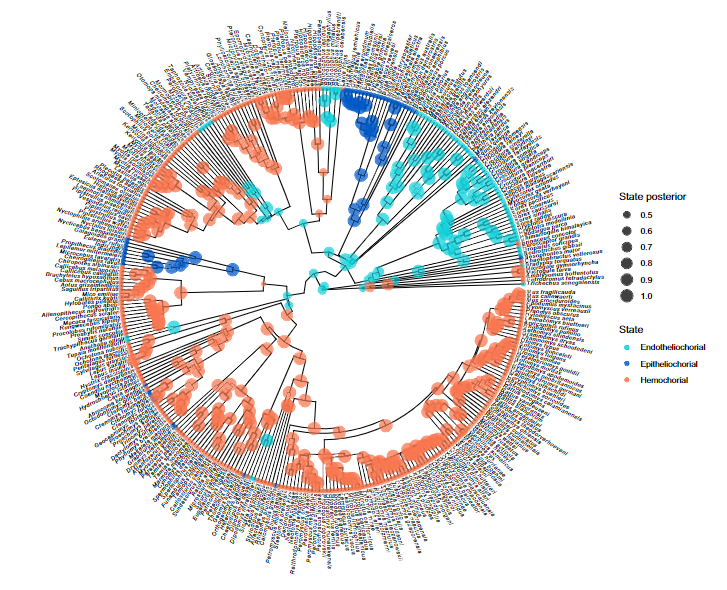
RevGadgets Statistical phylogenetic methods are the foundation for a wide range of evolutionary and epidemiological studies. However, as these methods grow increasingly complex, users often encounter significant challenges with summarising, visualising and communicating their key results. Here, Tribble et al. present RevGadgets, an R package for creating publication-quality figures from the results of a large variety of phylogenetic analyses performed in RevBayes.
Bulk arthropod abundance, biomass and diversity estimation Arthropod abundance, biomass and taxonomic diversity are key metrics often used to assess the efficacy of restoration efforts. Gathering these metrics is a slow and laborious process, quantified by an expert manually sorting and weighing arthropod specimens. Here, Schneider et al. present a tool to accelerate bulk arthropod classification and biomass estimates utilising machine learning methods for computer vision. The method simultaneously classifies >1,000 arthropods to functional groupings while estimating total and class specific biomass, without any computer vision bounding box or mask annotations, all from a single photo.
A simulation study on the importance of modelling the phylogeny Meta-analyses in ecology and evolution require special attention due to certain study characteristics in these fields. Primary articles in these fields usually report results that are observed from studies conducted with different species, and the phylogeny among the species violates the independence assumption. Also, articles frequently allow the computation of multiple effect sizes which cannot be accounted for by conventional meta-analytic models. While both issues can be dealt with by utilising a multilevel model that accounts for phylogeny, the performance of such a model has not yet been examined extensively. In this article, Cinar et al. investigate the performance of this model in comparison with simpler models.
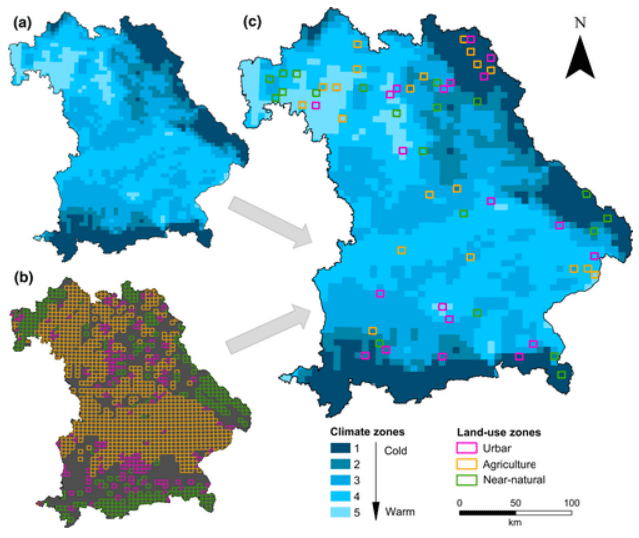
Disentangling effects of climate and land use on biodiversity and ecosystem services Climate and land-use change are key drivers of environmental degradation in the Anthropocene, but little is known about their interactive effects on biodiversity and ecosystem services. Long-term data on biodiversity trends are currently lacking. Furthermore, previous ecological studies have rarely considered climate and land use in a joint design, did not achieve variable independence or lost statistical power by not covering the full range of environmental gradients. Here, Redlich et al. introduce a multi-scale space-for-time study design to disentangle effects of climate and land use on biodiversity and ecosystem services.
Other Applications Articles
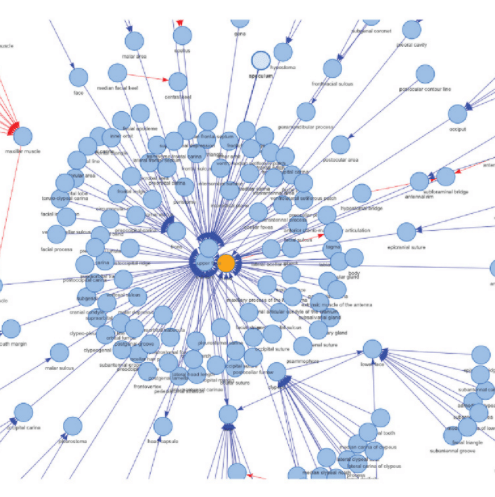
ontoFAST Ontologies are becoming a fundamental technology for analysing phenotypic data. The commonly used Entity–Quality (EQ) provides rich semantics for annotating phenotypes and characters using ontologie, but EQ syntax might be time inefficient if this granularity is unnecessary for downstream analysis. Here, Tarasov et al. present ontoFAST, an R package that aids fast annotations of characters with biological ontologies. ontoFAST takes a biomedical ontology in OBO format and a list of characters as input, and produces a list of mappings from characters to ontology terms as output.
allodb Allometric equations for calculation of tree above-ground biomass (AGB) form the basis for estimates of forest carbon storage and exchange with the atmosphere. While standard models exist to calculate forest biomass across the tropics, we lack a standardized tool for computing AGB across boreal and temperate regions that comprise the global extratropics. In answer to this, Gonzalez-Akre et al. present allodb, an integrated R package containing systematically-selected published allometric equations and proposed functions to compute AGB. The data component of the package is based on 701 woody species identified at 24 large Forest Global Earth Observatory (ForestGEO) forest dynamics plots representing a wide diversity of extratropical forests.
Other Practical Tools Articles
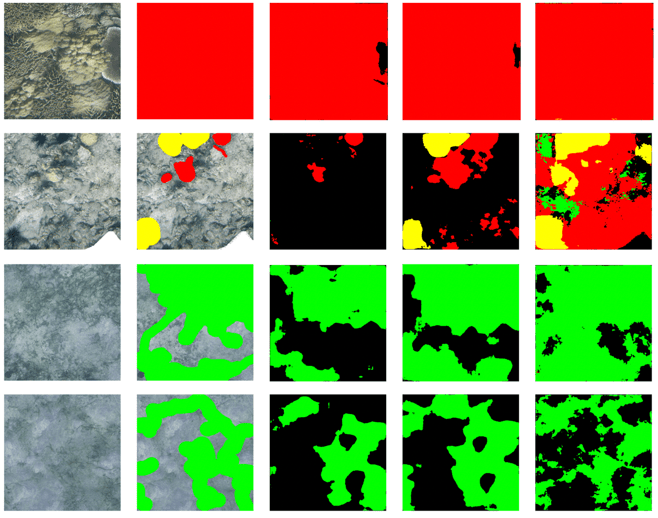
Cost-effective seafloor habitat mapping Various sampling and monitoring strategies have been developed to assess marine habitats and life-forms, but such methods remain expensive. In this article, Terayama et al. present a practical seafloor habitat mapping method combining a cost-effective survey system (P-SSS: portable speedy sea scanner) and a deep learning-based quantification method. The authors also present a segmentation method utilising the U-Net architecture to estimate the coverage of corals, seagrass and sea urchins using a large-scale 2D image. This highly portable survey system is expected to become a promising tool for marine environmental surveys.
The Storks on the Cover
Drs. Inge Muller and Wolfgang Fiedler check on the performance of tracking tags they fit to 20 young storks in Poland. As the birds fledged and began migrating, they flew beyond the range of the biologist’s antenna, but their tags connected to phone networks along the way, streaming live data into the Movebank system, presented by Kays et al. in their article. Not all young are successful in their first migration from Europe, through the Middle East, and into Africa. Drs. Muller and Fiedler watch for a lack of movement or drop in temperature suggesting a potential problem, and then investigate themselves if they can. With the addition of the Animal Tracker App, it became possible to study bird fate globally by asking for help from collaborators or citizen scientists. The locals could see the location of the tag, and report back to the scientists the bird’s fate, which included cases of electrocution on powerlines, hunting, predation, and illness. In a few cases, citizens got to the bird in time to rescue it from being stuck in some kind of human infrastructure. Eventually, successful migrants return to their breeding grounds, where tracking data show nesting behaviour, and the start of a new generation of storks. Photo credit: ©Christian Ziegler, MPIAB.