Our July Issue is now online! This issue contains 20 articles about the latest methods in ecology and evolution, including methods for characterising soil bacterial biodiversity, identifying fish species in fish markets using eDNA, standarising and cleaning biodiversity data and much more! Plus, read the editorial about our switch to a gold open access model this month. Read on to find out about this month’s featured articles and the article behind our fabulous duck cover.
Editorial: An Open Future for MEE
The shift towards open access publishing of research has been occurring for the past two decades and has accelerated recently with some of the world’s largest research funders mandating that their funded research should soon be published only in fully open access journals. The time has now come for MEE to take the next step and become a fully open access journal. From July 6, all articles submitted to us will ultimately be published open access, and all existing content will become open access from January 2023. Read the full editorial to find out more.
Featured Articles
An analysis of simulating algorithms and graph metrics used to quantify topology River networks are frequently simulated for use in the development and testing of ecological theory. Currently, two main algorithms are used, stochastic branching networks (SBNs) and optimal channel networks (OCNs). The topology of these simulated networks and ‘real’ rivers is often quantified using graph theoretic metrics; however, to date, there has not been a comprehensive analysis of how these algorithms compare regarding graph theoretic metrics, or an analysis of metric redundancy and variability across dendritic ecological networks. Lee et al. aim to provide guidance as to which algorithm and metrics should be used, and under what circumstances.
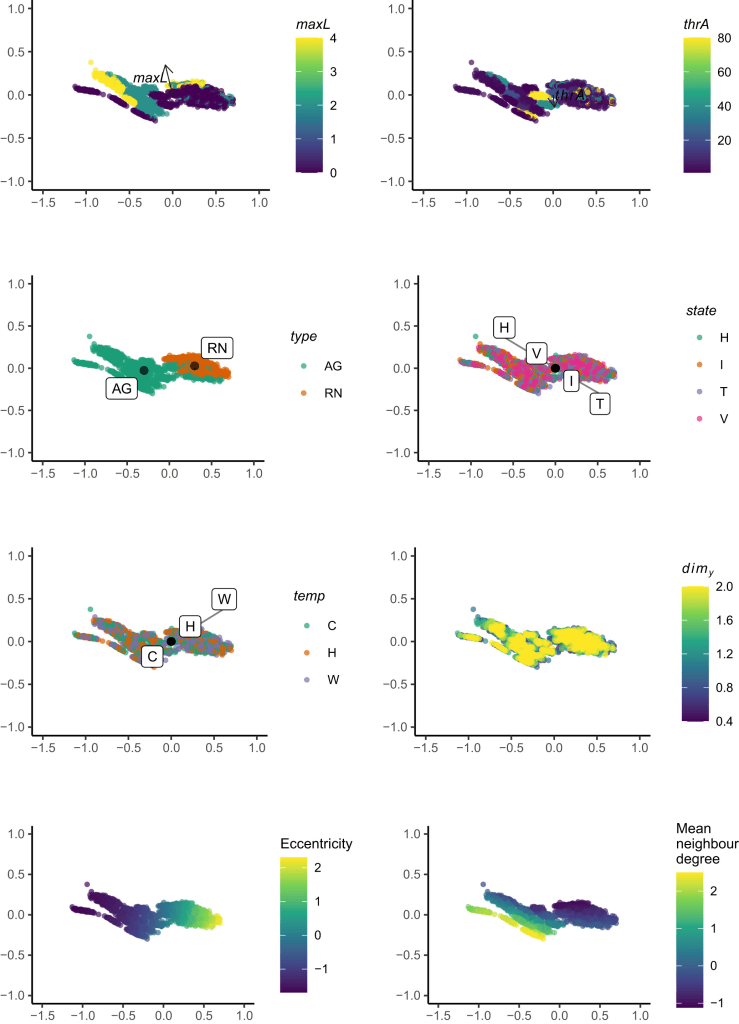
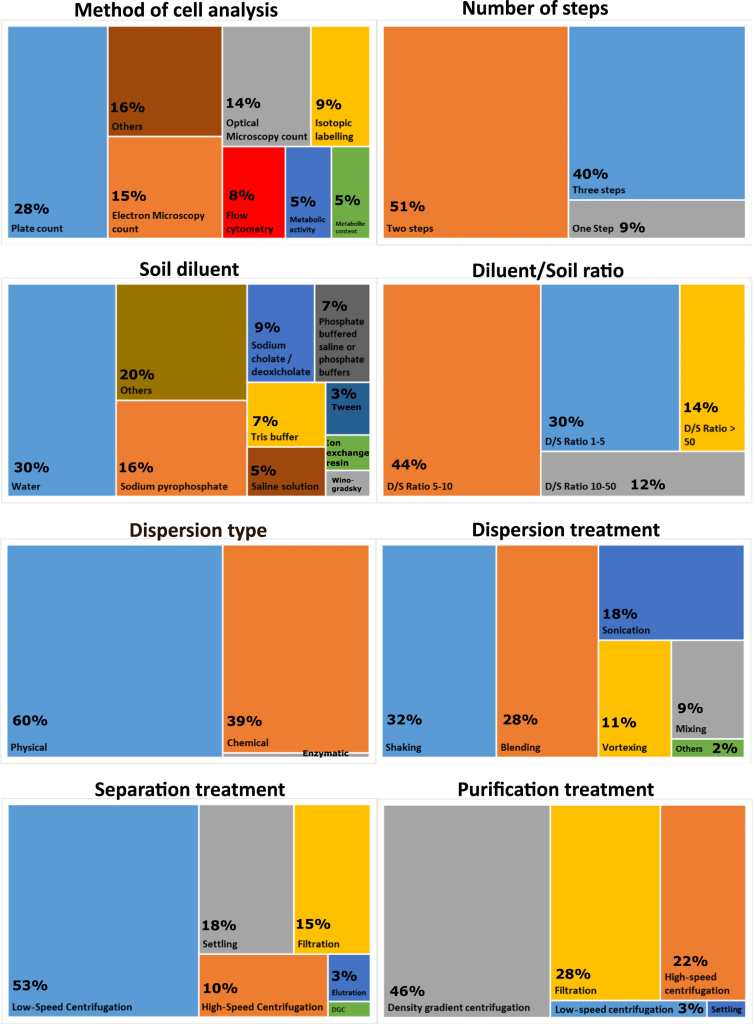
Soil bacterial biodiversity characterization by flow cytometry Flow cytometry (FCM) is a powerful technique to characterize cell communities due to its high robustness and accuracy, requiring only a short time for the characterization. Therefore, FCM could expand soil research capabilities by allowing the characterization of different aspects of bacterial biodiversity. Here, El Mujtar et al. review the state-of-art of the use of flow cytometry in scientific research and the protocols used for the extraction of bacteria from soil. They analyse the literature to take stock of the diversity of methodologies for soil processing and applications of flow cytometry in bacterial characterization considering abundance, diversity, community structure and functional properties.
A toolkit for standardizing, integrating and cleaning biodiversity data The increase in online and openly accessible biodiversity databases provides a vast and invaluable resource to support research and policy. However, without scrutiny, errors in primary species occurrence data can lead to erroneous results and misleading information. Here, Ribeiro et al. introduce the Biodiversity Data Cleaning (bdc), an R package to address quality issues and improve the fitness-for-use of biodiversity datasets. The bdc package brings together several aspects of biodiversity data cleaning in one place.
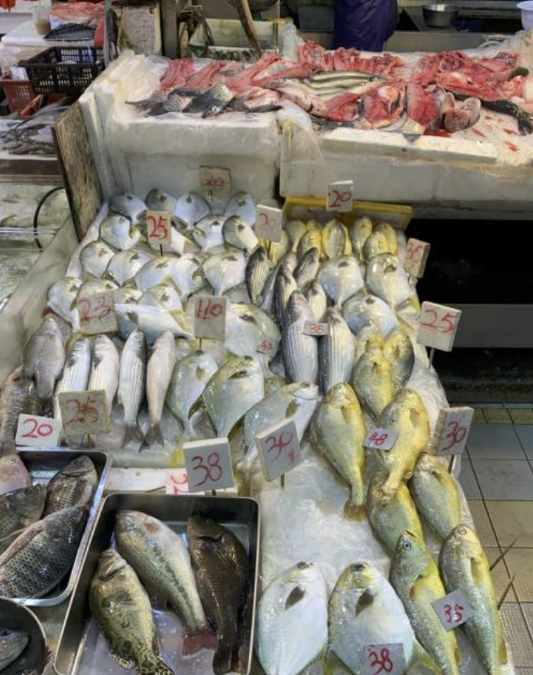
eDNA-based survey method for urban fish markets The state of knowledge for many fish species is limited by current monitoring techniques, which rely on labour-intensive visual or genetic surveys of individual specimens (often at inconsistent or coarse taxonomic resolution). To address the need for more efficient methods that effectively monitor trade, Richards et al. have developed a novel application of eDNA-based metabarcoding that can identify a broad range of fish taxa from effluent water draining from urban fish markets.
Synthetic microbial consortia with programmable ecological interactions
Central to the composition, structure and function of any microbial community is the complex species interaction web. But understanding the overwhelming complexity of ecological interaction webs has been challenging, owing at least partly to the lack of efficient tools for disentangling species interactions in natural or artificial microbial communities. In this study, Li et al. have developed a microbial experimental system which allows for rapidly generating microbial consortia with programmable ecological interactions.
The Duck on the Cover
A tracker-equipped Pacific Black Duck Anas superciliosa flapping its wings while standing on a log in a farm dam in rural Victoria, Australia. Aside from GPS positions, the tracker developed by Druid Technology is able to record and translate raw accelerometer data into behaviours every 2 seconds, which are stored on board and are downloaded using either 3G or Bluetooth. The translation of accelerometer data to behaviour is performed by an algorithm remotely uploaded to the tracker. This algorithm is developed using machine learning on simultaneously collected behavioral observations and accelerometer data sent to the observer via Bluetooth. On-board processing and data transmission proved energy efficient and came at a low weight cost to the tracker, providing great potential to open up new areas in ecological and behavioural research on a wide array of taxa. Image credit: ©Trent Leen.