(this is the first in a possibly irregular series of posts about papers that catch my eye. I don’t intend to only cover MEE papers, but I had to start somewhere)
![]() A perennial worry for anyone building models for the real world is whether they actually represent the real world. If the whole process of finding and fitting a model has been done well, the model will represent the data. But the data is only part of the real world. How can we be sure our model will extrapolated beyond the data?
A perennial worry for anyone building models for the real world is whether they actually represent the real world. If the whole process of finding and fitting a model has been done well, the model will represent the data. But the data is only part of the real world. How can we be sure our model will extrapolated beyond the data?
If we have some extra data we can check this by seeing if the model fits well to the new data. If we don’t, then we can cheat by splitting out data into two sets: we use one to fit the model and a second to see how well the model fits to new data. This approach is used a lot, for example when fitting species distribution models (SDM), but it’s not really correct. The problem is that the data were all collected at the same time, so any peculiarities to the data (e.g. because it was collectd in a single year) will remain. A model that fits well to the data might not fit to next year’s data. We talk about this as the model over-fitting: it fits to the peculiarities of the data, rather than summarising the underlying biology.
Over the last few years there has been a lot of species distribution modelling going on. Most of this involves throwing the data into a black box along with some data from WorldClim and seeing what it spits out. Inside a lot of these black boxes use machine learning methods like neural networks and random forests. One of the reasons for using these methods is that they perform well under cross-validation: i.e. when 30% (say) of the data are randomly removed from the data in the fitting of the model, and then the model is used to predict these data. But these data are correlated with the data that is used in the ftting, so this is not an independent test of the model. So, how well do these methods work with independent data?
Well, guess what? Someone has checked. In the most recent issue of MEE, Seth Wenger of Trout Unlimited and Julian Olden of the University of Washington report their findings. Their data is the distribution of brook and brown trout in the western US. What they do is to fit distribution models to their data, and try two types of cross-valudation: first the traditional sort, removing data at random, and second by removing data in bands:
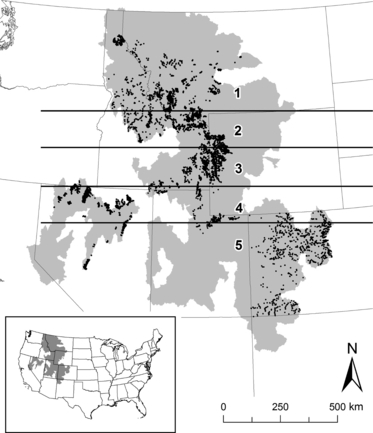
They then compared the measures of how well the models did. With the standard cross-valudation, random forests did best, followed by neural nets, and standard generalized linear mixed models (GLMMs, i.e. fitting straight lines and quadratic curves) were the the worst. But when the fitted models were used to predict the data in the bands, the situation was reverse: the GLMMs did best.
This suggests that random forests and neural nets are over-fitting the data: the reason they do so well is that the data that is used to test the model is too close to the data used to fit it. Another reason to think that they are over-fitting is that the fitted curves just don’t look sensible:
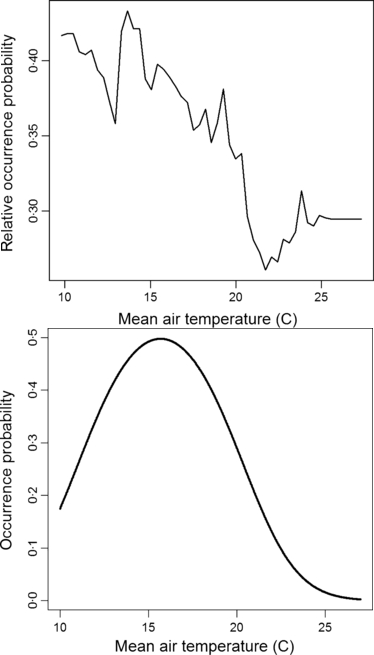
Do you really believe such a complicated curve for random forests (at the top)? This is not peculiar to this study: I’ve seen horrible plots like this in other studies (MAXENT also produces uninterpretable curves).
So, what’s the take-home message? Just that simpler models seem to do better than machine learning models, which end up being just too damn complicated. I like this result, largely because I generally use these simpler models, but also because the simpler models are easier to understand. Anything that tells us we can make life simpler is always attractive.
Reference
Wenger, S., & Olden, J. (2012). Assessing transferability of ecological models: an underappreciated aspect of statistical validation Methods in Ecology and Evolution, 3 (2), 260-267 DOI: 10.1111/j.2041-210X.2011.00170.x
Relatesd to this (and I only saw it after I had written this post) is this new paper in Ecology, which looks at the effects of spatial bias in the presences used in presence-only methods. I haven’t read it thoroughly, but it’s pointing to the same problem.
Nice post Bob, and thanks for the link to the Ecology paper. I haven’t got access to the MME paper (must email librarian) but both response curves could potentially be wrong. The RF one suggests the optima for the species is not within the range of the data and you could approximate the RF curve with a sigmoid fit. So is the GLMM over-specified too? Without seeing the data it is hard to comment further, but the estimated “curves” at the low end of the temperature gradient are so very different it does somewhat beg the question what is the response down that end?
I would have liked to have seen a GAMM as well: I find quadratics are generally too stiff. It’s possible there’s very little data at the lower end but the probability should plateau, and of course the quadratic has to decline somewhere.
That reminds me of a paper I must get back to, Bob! And I am increasingly wondering if the first port of call for our work should be an ecology journal rather than a statistical journal…
At the beginning of the discussion you seem to be discussing temporal and spatial scales that may affect the process under study, suggesting that partitioning the data in two sets may not be appropriate. However, then you go on discussing a paper that seems to be be doing the same thing: spatial partition (bands). It seems to me that the first part of the post is a distraction from the main thrust of the discussion that is model overfitting or, even, learning vs statistical models.
I don’t have access to the paper right now, but I’d expect that, in general, a model with some theoretical underpinnings on the actual biological problem (or at least that reflects biological constraints) would over perform those that optimize local fit. We often see that in applied forestry models.
The problem is how the partitioning is done – standard cross-validation draws the fitting data randomly from the whole data. The paper asks what happens if we split by geographic area, and shows a different result.
I 100% agree that using a biologically realistic model sould be better, I’m co-editor of a special issue that should be coming out in the Journal of Biogeography this year where we’re asking how to do this.
I really that the problem is not using complicated models per se, but blindly using them without (1) doing some simple exploratory analysis first to understand the data and get to know its limitations and peculiarities, (2) understanding the limitations of those models, (3). trying to stablish if this data even needs a complicated model or can be reasonably understood under simpler models.
If I’m modelling the effect of different conditions over some variable I’m interested in, and the first thing to do is a freaking linear factor model! AFTER that I may conclude it wasn’t enough, the data have too complicated a structure for simple linear models to capture or whatever. But hey… I might have saved myself some work and I certainly will get to know more about the data in the process so that I may choose the next model with more confidence.
I usually see people just fitting whatever models they have black box software that works in a few minutes available. Those guys hardly understand what are the underlying assumptions, the limitations and the peculiarities of each algorithm.