Alessandro Filazzola & Christopher Lortie tell us about their new article ‘A call for clean code to effectively communicate science‘, which provides a series of recommendations and a suite of tools that can be used to help support scientists to produce cleaner code.
Can you clearly understand the code that you have written? What about if you gave it to a colleague? Or a reviewer? Or even a future version of yourself? Writing code that is as legible as it is functional is not an easy task, but it has significant benefits. Code that is understandable is less likely to have errors, more easily reviewed by colleagues, and more easily replicated. Thus, computational reproducibility (i.e., the ability to recreate someone’s analysis and results) is just as important as experimental reproducibility. That is why many journals, Methods in Ecology and Evolution included, encourage code publishing with manuscripts. However, what good is that published code if it is illegible?
Alessandro Filazzola and Christopher Lortie are both field ecologists and computational biologists. Neither is trained as a computer programmer and each entered the field of ecology through surveying the natural environment. Overtime, reading books on R, coding, and statistics, it became apparent that there are a set of best practices that should be followed when coding in ecology and evolution. One of the more influential reads was the book by Robert Martin called Clean Code. While this book was written for a software developer using Java, the examples were easily to follow along because the code was well written. That is the ultimate goal of clean code – having the code understandable by someone who is not familiar with the language.
What is clean code?
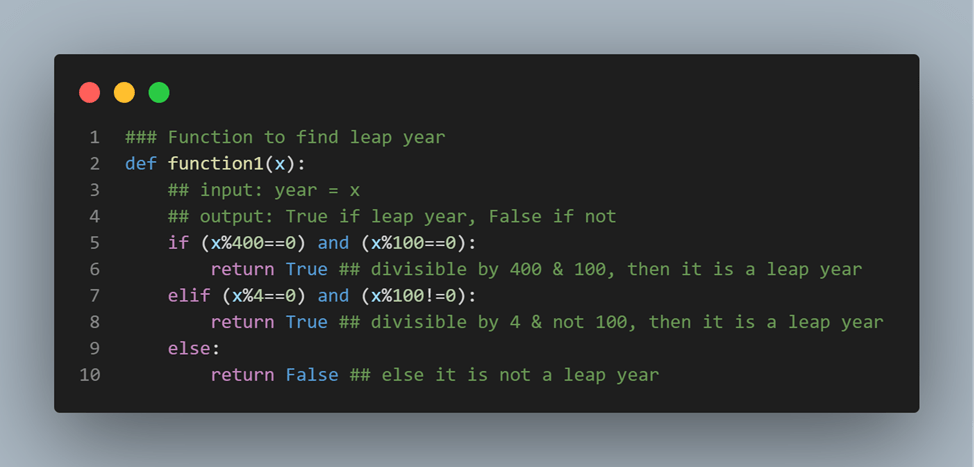

Let’s look at an example of how code can be improved to be more easily understandable. In the above example, we have two chunks in Python that do a simple enough task of checking for leap years. However, there are certainly a few differences between these two chunks. In the first, the user does not use clearly understandable function names or parameters. Instead, they rely on the comments to provide that additional clarity. By providing more explicit function or parameter names, while longer to write, can save the need for writing lengthy comments. Additionally, most IntelliSense in integrated development environments (IDEs) such as RStudio, PyCharm, or VSCode, have autocomplete for longer function names. Additionally, these two examples show differences in spacing around operators and text. Spacing can be a powerful tool in improving legibility just the same as it is when writing in English or other languages.
Best practices for clean code
In the article A call for clean code to effectively communicate science, authors Filazzola and Lortie detail the incentives to write clean code, the basics when crafting, and the implications for the typical computational biologist. The authors provide a table of best practices that includes formatting styles such as indenting, horizontal spacing, naming conventions, vertical ordering, among many others. At its core, clean code should be able to communicate effectively, be formatted for easy legibility, and include abstraction to focus the user on more important concepts.
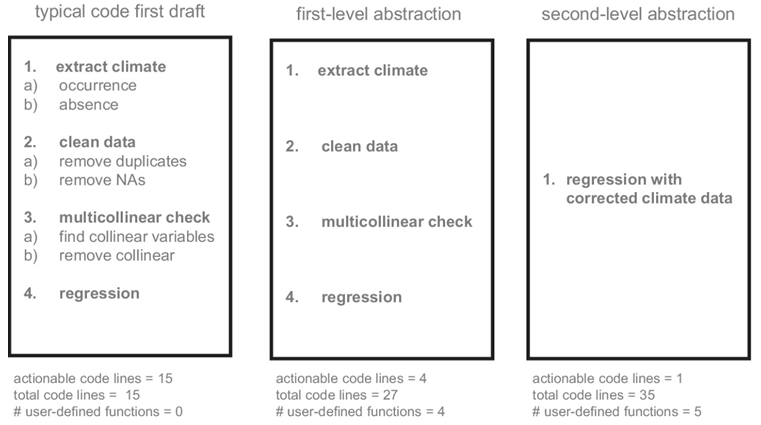
The idea of abstraction is one of the more complicated topics introduced that borrows from computer science. In short, abstraction involves removing complex elements from the executable parts of the code to focus the users on the chunks that require greater attention. For instance, think about the `lm` function in R to conduct linear models, which itself is a type of abstraction. When we use the `lm` function we are not concerned with the code that calculates the sum of squares, estimates of the regression coefficients, or converts F-statistics into p-values. We are generally only concerned with the predictors, response variable, and sometimes a few additional parameters. The `lm` function is an abstraction of a much larger code base that allows us, the users, to focus more on the sections of the script that need greater attention. Similarly, we can apply abstraction to our code by wrapping sections of code into functions to increase focus. In the paper, the authors detail an example of simplifying an R script by wrapping chunks into functions to reduce the number of lines that are actionable by the user (see example below).
Using tools to help when writing clean code
Writing clean code can be hard, but it does not have to be. There are many tools available that can support writing clean code. Many IDEs have extensions or built-in functionality that accomplish many of the best practices identified in the manuscript, such as automatic formatting (i.e., linters). There are also style guides providing by tech companies and language specific developers. The British Ecological Society produced a coding style guide in 2017. A call for clean code to effectively communicate science not only provides a series of recommendations, but also the suite of tools that could be used to help support us as scientists produce cleaner code. While we did not get into this field to become programmers, with coding becoming increasingly integrated into our daily work, we can certainly adopt some best practices to craft code that is legible and reproducible.
Just read and follow the 1974 book The Elements of Programming Style by Kernighan and Plauger. They had this concept dialed decades ago.
Irrelevant but:
all numbers divisible by 400 are also divisible by 100, so the first condition can just be
if year % 400:
return true