Individual History and the Matrix Projection Model
Post provided by Rich Shefferson
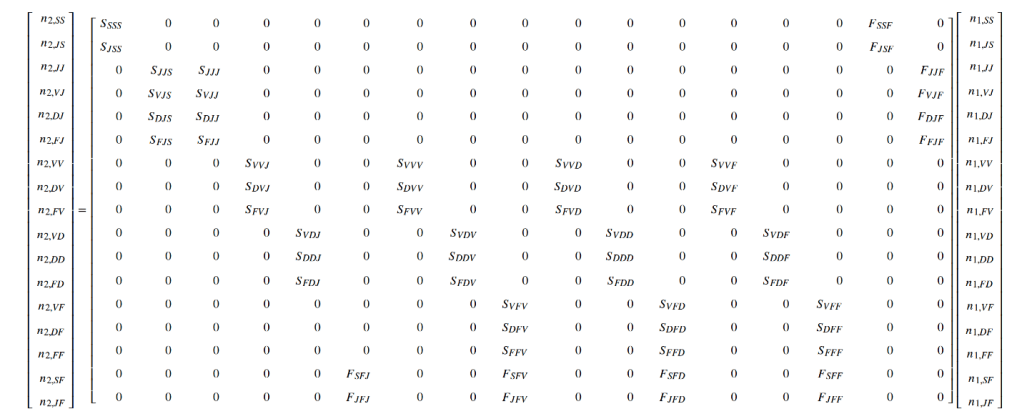
Matrix projection modeling is a mainstay of population ecology. Ecologists working in natural area management and conservation, as well as in theoretical and academic realms such as the study of life history evolution, develop and use these models routinely. Matrix projection models (MPMs) have advanced dramatically in complexity over the years, originating from age-based and stage-based matrix models parameterized directly from the data, to complex matrices developed from statistical models of vital rates such as integral projection models (IPMs) and age-by-stage models. We consider IPMs to be a class of function-based MPM, while age-by-stage MPMs may be raw or function-based, but are typically the latter due to a better ability to handle smaller dataset. The rapid development of these methods can leave many feeling bewildered if they need to use these methods but lack sufficient understanding of scientific programming and of the background theory to analyze them properly.
Continue reading “Individual History and the Matrix Projection Model”






