Post provided by Jordan Cuff, Fredric Windsor, Maximillian Tercel, James Kitson and Darren Evans.
Do you spend your days incapacitated by the agony of not knowing how to combine your field-based observational data with your dietary metabarcoding results? Perhaps every time you go to merge them, the horror of conflicting data types and biases causes you to run and hide from the mere thought of analysis. Or maybe the thought of such problems hadn’t even crossed your mind! In this post, Jordan Cuff and co-authors share insight from their recent publication on using dietary metabarcoding in network ecology and how to merge metabarcoding with traditional data types.
A blossoming relationship between dietary metabarcoding and networks
The advent of metabarcoding has been a paradigm shift in dietary analysis, allowing us to study the diet of even cryptic, tiny, or fluid-feeding animals. Dietary metabarcoding can be used to detect interactions between species by identifying the resources in a consumer’s gut and faeces using the DNA of that resource species. This is an incredibly sensitive method that can detect minute quantities of consumed resources, including soft-bodied, tiny and morphologically cryptic species often neglected by traditional methods.
Dietary metabarcoding hasn’t yet been used widely for trophic interactions in a network context. There are a few cool examples examining the diets of bats and spiders, as well as exploring ecological networks in Costa Rica and the Mediterranean, but it’s still very much a growing subfield. As networks become progressively complex and ambitious (especially multilayer ones), dietary metabarcoding is increasingly important. That being said, there are some important things to consider, particularly when merging interaction data from metabarcoding with data from other sources.
Quantitative metabarcoding: counting with a broken abacus
Quantification is a topic that can divide a room and cause arguments between otherwise collegial metabarcoders. Some believe that the number of sequencing reads from a sample reflects the abundance of the corresponding species in the sample, but others believe that these values are wildly inaccurate due to PCR bias among other factors. Whatever side of the debate you’re on, no one can deny that quantification and metabarcoding have a tricky relationship.
On one hand, if you use the number of sequencing reads from metabarcoding to quantify how much of something an animal has eaten, it can introduce bias. On the other hand, if you convert these data to binary presence/absence values, you neglect the fact that things will be eaten in differing proportions altogether. Some researchers use frequency-of-occurrence data (counts of how many of each organism in a group eat a certain species) to provide some level of quantification from otherwise binary data. But if these aren’t normalised (all values for each individual made to add up to one before combining within a group) the data are still skewed, particularly between generalists and specialists.
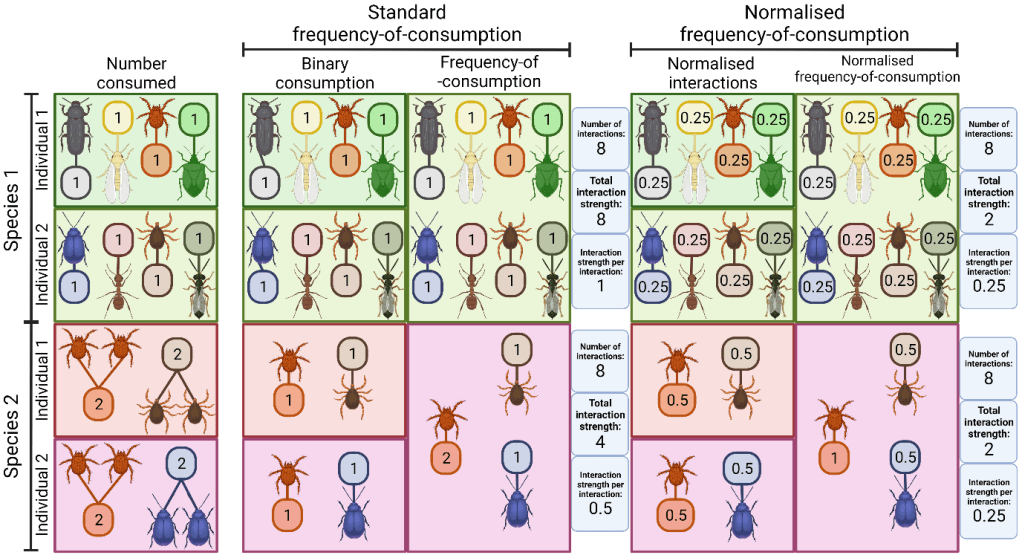
For example, in the image above the two species are eating the same number of individuals. Species 1 is a generalist with the two individuals each eating four different prey species. Species 2 is a specialist with the two individuals eating only two prey species each, one of which overlaps, but they still eat four prey individuals each. If these are added up from binary data and frequency-of-occurrence calculated, species 1 has a total interaction strength (the number of interactions included in the network) of 8, whereas species 2 has a total interaction strength of 4, despite both species having the same number of interactions. By first normalising these values through including each eaten species as a fraction of one for each individual predator, the total interaction strength is equal between the two species.
It’s all a matter of perspective
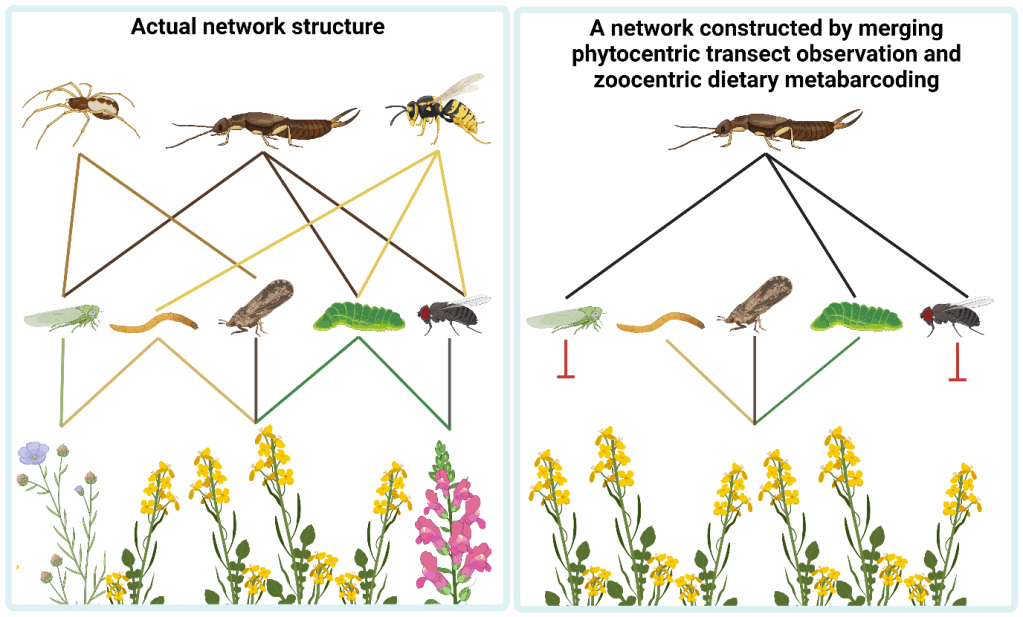
Metabarcoding and traditional transect surveys also differ in their perspective. Transects might be phytocentric (plant-based, with many interacting species) whereas dietary metabarcoding is zoocentric (animal-based, with many consumed species), which can have some tricky implications. For example, the zoocentric perspective of dietary metabarcoding reduces diversity at the top layer of a network, whereas phytocentric observations of herbivory reduces diversity at the bottom layer. If these two datasets are merged to construct a tripartite network, the top and bottom layers are reduced, leaving a bloated middle layer. This is why we recommend trying to sample different network layers from both perspectives where possible. Keeping a keen eye on sampling completeness for each method is also key!
Biases, errors and misidentifications
Metabarcoding is taxonomically biased depending on the similarity in sequence between PCR primers and target DNA. This means that some interactions might be left out of the network, reducing the overall interaction diversity. We suggest using multiple complementary primer pairs together to represent different overlapping compartments of the full breadth of consumed species. These primers can be tested in silico (via software that simulates PCR), in vitro (by testing the primers against DNA extracts in the lab) and through mock community assessment (by looking at how well represented a known mixture of species is after sequencing) to predict how these biases might affect the data.
False positives are also a massive issue in metabarcoding data. Contaminants from the environment or lab, or even errors produced during sequencing, can result in incorrect conclusions about the diet of animals with sometimes quite obscure consequences (e.g., tiny money spiders eating people on farms). Researchers have often used “minimum sequence copy thresholds” (i.e., if read counts aren’t above a set value, they are removed) to get rid of these contaminants, but these can often remove legitimate data. We therefore suggest thoughtful and measured use of minimum sequence copy thresholds. Poor reference databases can also reduce the quality of data produced by metabarcoding, so checking the completeness of databases is crucial before embarking on the metabarcoding journey. Creating your own tailor-made database is ideal, but expensive and laborious for many taxa.
Accidents and omnivores
Accidental and secondary consumption are also rife in metabarcoding, but their importance depends on the questions being asked. Does it matter that a beetle is accidentally eating a parasitoid inside the body of an aphid, or that a spider is ingesting barely any DNA from the gut contents of its prey? Perhaps not in some circumstances, but, if interpreted without a healthy dose of cynicism these data could be responsible for turning a biocontrol agent into a pest, for example. This is particularly problematic for omnivores, especially when only looking at the plant component of their diet. More than this, secondary consumption might lead you to think that a carnivore is an omnivore!

Dietary metabarcoding isn’t without its problems, but it provides a really powerful tool for constructing increasingly complex networks, particularly including species that have traditionally been left out of analysis. While dietary metabarcoding may revolutionise our ability to construct these complex ecological networks, we need to use it with caution. If done right, we could be building even bigger and better networks. By considering the integration of additional data types, perhaps even nutrients, networks can become more informative and address increasingly ambitious ecological questions. Hopefully, our article helps to identify the problems involved in merging these data types before they appear in the data.
Follow the link below to read the full Methods in Ecology and Evolution article:
Overcoming the pitfalls of merging dietary metabarcoding into ecological networks

One thought on “Merging dietary metabarcoding into networks: turning “don’ts” into “dos””